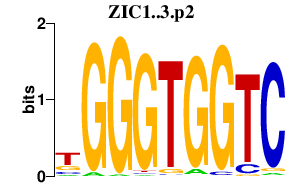
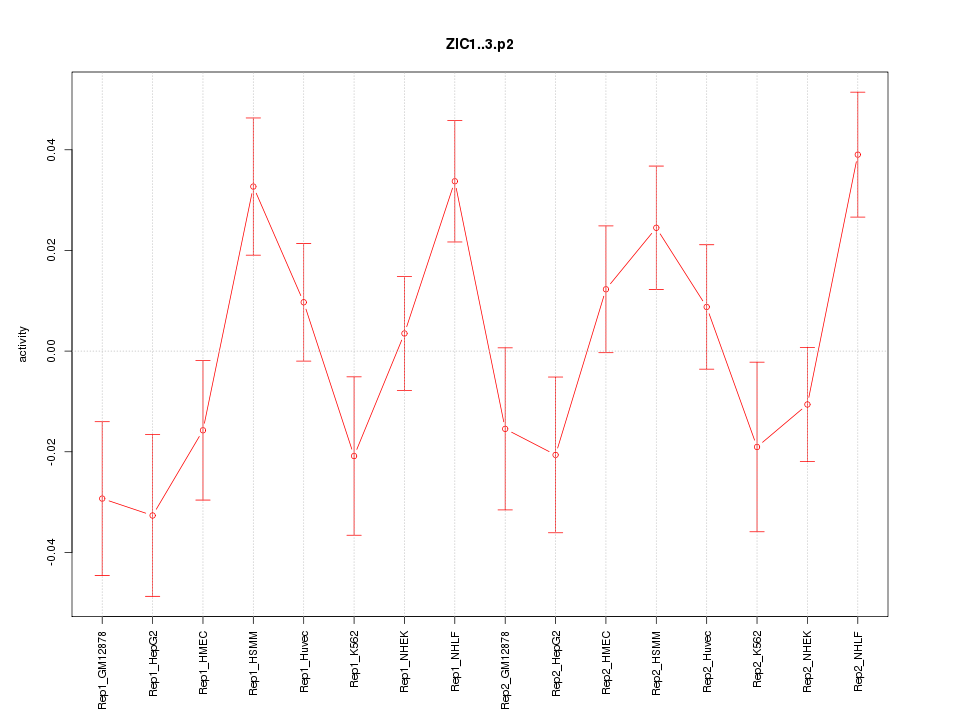
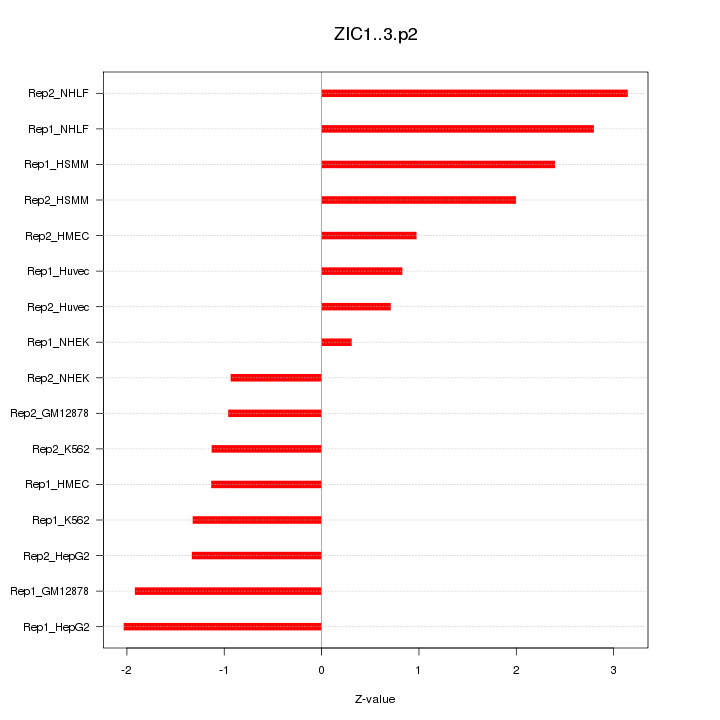
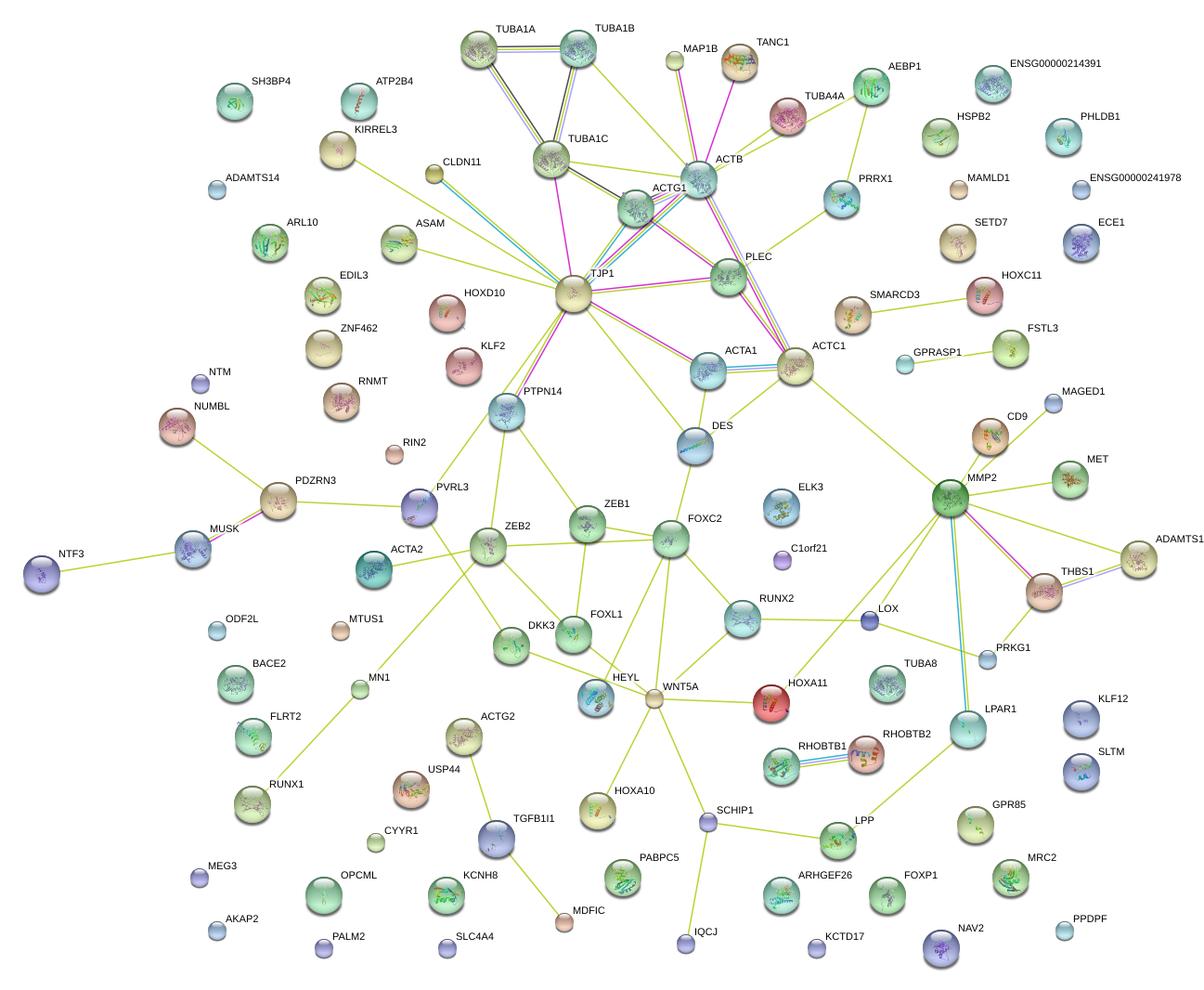
|
chr16_+_85169615
|
4.596
|
NM_005250
|
FOXL1
|
forkhead box L1
|
|
chr3_+_160964574
|
4.360
|
|
SCHIP1
|
schwannomin interacting protein 1
|
|
chr3_-_73756668
|
4.081
|
NM_015009
|
PDZRN3
|
PDZ domain containing ring finger 3
|
|
chr15_+_37660568
|
3.467
|
NM_003246
|
THBS1
|
thrombospondin 1
|
|
chr4_+_72271218
|
3.167
|
NM_001098484
NM_001134742
|
SLC4A4
|
solute carrier family 4, sodium bicarbonate cotransporter, member 4
|
|
chr3_+_64645819
|
2.893
|
|
LOC100507098
|
hypothetical LOC100507098
|
|
chr9_-_112840064
|
2.441
|
NM_001401
NM_057159
|
LPAR1
|
lysophosphatidic acid receptor 1
|
|
chr21_-_27139143
|
2.308
|
|
ADAMTS1
|
ADAM metallopeptidase with thrombospondin type 1 motif, 1
|
|
chr10_+_52421120
|
2.306
|
|
PRKG1
|
protein kinase, cGMP-dependent, type I
|
|
chr16_+_85158357
|
2.285
|
NM_005251
|
FOXC2
|
forkhead box C2 (MFH-1, mesenchyme forkhead 1)
|
|
chr21_-_27139563
|
2.195
|
NM_006988
|
ADAMTS1
|
ADAM metallopeptidase with thrombospondin type 1 motif, 1
|
|
chr11_+_19691397
|
2.147
|
NM_145117
NM_182964
|
NAV2
|
neuron navigator 2
|
|
chr9_+_111850698
|
2.139
|
NM_001004065
NM_001198656
|
AKAP2
|
A kinase (PRKA) anchor protein 2
|
|
chr9_+_108665168
|
2.133
|
NM_021224
|
ZNF462
|
zinc finger protein 462
|
|
chr20_+_19903778
|
2.129
|
|
RIN2
|
Ras and Rab interactor 2
|
|
chr7_-_27191287
|
2.034
|
NM_005523
|
HOXA11
|
homeobox A11
|
|
chr14_+_100362236
|
2.005
|
|
MEG3
|
maternally expressed 3 (non-protein coding)
|
|
chr3_+_155321826
|
1.970
|
NM_015595
|
ARHGEF26
|
Rho guanine nucleotide exchange factor (GEF) 26
|
|
chr14_+_100362204
|
1.898
|
|
MEG3
|
maternally expressed 3 (non-protein coding)
|
|
chr16_+_54070302
|
1.891
|
NM_004530
|
MMP2
|
matrix metallopeptidase 2 (gelatinase A, 72kDa gelatinase, 72kDa type IV collagenase)
|
|
chr11_+_111288686
|
1.845
|
|
HSPB2-C11ORF52
HSPB2
|
HSPB2-C11orf52 read-through transcript
heat shock 27kDa protein 2
|
|
chr2_-_144991386
|
1.705
|
|
ZEB2
|
zinc finger E-box binding homeobox 2
|
|
chr2_-_144991556
|
1.666
|
|
ZEB2
|
zinc finger E-box binding homeobox 2
|
|
chr5_-_121440704
|
1.662
|
NM_001178102
|
LOX
|
lysyl oxidase
|
|
chr22_-_26527469
|
1.622
|
NM_002430
|
MN1
|
meningioma (disrupted in balanced translocation) 1
|
|
chr11_+_111288669
|
1.560
|
NM_001541
|
HSPB2-C11ORF52
HSPB2
|
HSPB2-C11orf52 read-through transcript
heat shock 27kDa protein 2
|
|
chrX_+_90576252
|
1.545
|
NM_080832
|
PABPC5
|
poly(A) binding protein, cytoplasmic 5
|
|
chr11_-_126375864
|
1.538
|
|
KIRREL3
|
kin of IRRE like 3 (Drosophila)
|
|
chr14_+_85066265
|
1.535
|
|
FLRT2
|
fibronectin leucine rich transmembrane protein 2
|
|
chr19_-_45888259
|
1.528
|
NM_004756
|
NUMBL
|
numb homolog (Drosophila)-like
|
|
chr4_-_140696740
|
1.516
|
NM_030648
|
SETD7
|
SET domain containing (lysine methyltransferase) 7
|
|
chr11_+_131285747
|
1.512
|
NM_001144058
NM_001144059
NM_016522
|
NTM
|
neurotrimin
|
|
chr7_-_106088565
|
1.508
|
NM_175884
|
FLJ36031
|
hypothetical protein FLJ36031
|
|
chr17_+_58058783
|
1.495
|
|
MRC2
|
mannose receptor, C type 2
|
|
chr3_-_71196696
|
1.489
|
|
FOXP1
|
forkhead box P1
|
|
chr1_-_21478571
|
1.487
|
NM_001113347
|
ECE1
|
endothelin converting enzyme 1
|
|
chr7_-_27180398
|
1.482
|
NM_018951
|
HOXA10
|
homeobox A10
|
|
chr10_-_62373828
|
1.479
|
NM_014836
|
RHOBTB1
|
Rho-related BTB domain containing 1
|
|
chr5_-_92982927
|
1.474
|
|
|
|
|
chr5_+_175725050
|
1.471
|
NM_173664
|
ARL10
|
ADP-ribosylation factor-like 10
|
|
chr7_-_150576634
|
1.443
|
|
SMARCD3
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 3
|
|
chr2_+_159533329
|
1.418
|
NM_001145909
NM_033394
|
TANC1
|
tetratricopeptide repeat, ankyrin repeat and coiled-coil containing 1
|
|
chr13_-_73605914
|
1.383
|
NM_007249
|
KLF12
|
Kruppel-like factor 12
|
|
chr1_+_168899936
|
1.378
|
NM_006902
NM_022716
|
PRRX1
|
paired related homeobox 1
|
|
chr22_+_35777561
|
1.360
|
NM_024681
|
KCTD17
|
potassium channel tetramerisation domain containing 17
|
|
chr6_+_45498354
|
1.341
|
|
RUNX2
|
runt-related transcription factor 2
|
|
chr15_-_27901828
|
1.339
|
NM_003257
NM_175610
|
TJP1
|
tight junction protein 1 (zona occludens 1)
|
|
chr3_+_189353502
|
1.335
|
|
LPP
|
LIM domain containing preferred translocation partner in lipoma
|
|
chr11_-_126375895
|
1.334
|
NM_001161707
NM_032531
|
KIRREL3
|
kin of IRRE like 3 (Drosophila)
|
|
chr19_+_627346
|
1.326
|
NM_005860
|
FSTL3
|
follistatin-like 3 (secreted glycoprotein)
|
|
chr3_-_55490401
|
1.316
|
|
WNT5A
|
wingless-type MMTV integration site family, member 5A
|
|
chr21_+_41461962
|
1.309
|
|
BACE2
|
beta-site APP-cleaving enzyme 2
|
|
chr8_-_17599438
|
1.297
|
NM_020749
|
MTUS1
|
microtubule associated tumor suppressor 1
|
|
chr11_-_11986704
|
1.292
|
NM_001018057
|
DKK3
|
dickkopf homolog 3 (Xenopus laevis)
|
|
chr7_+_44110527
|
1.290
|
|
AEBP1
|
AE binding protein 1
|
|
chr11_+_117983515
|
1.286
|
NM_001144758
NM_001144759
|
PHLDB1
|
pleckstrin homology-like domain, family B, member 1
|
|
chr6_+_45498262
|
1.278
|
|
RUNX2
|
runt-related transcription factor 2
|
|
chr1_-_212791188
|
1.266
|
|
PTPN14
|
protein tyrosine phosphatase, non-receptor type 14
|
|
chr3_+_112273352
|
1.252
|
|
PVRL3
|
poliovirus receptor-related 3
|
|
chr12_+_6179736
|
1.235
|
NM_001769
|
CD9
|
CD9 molecule
|
|
chr7_-_112513379
|
1.230
|
NM_001146266
NM_001146267
|
GPR85
|
G protein-coupled receptor 85
|
|
chr3_+_171619346
|
1.228
|
NM_005602
|
CLDN11
|
claudin 11
|
|
chr5_+_71438796
|
1.208
|
NM_005909
|
MAP1B
|
microtubule-associated protein 1B
|
|
chr7_-_150576660
|
1.198
|
NM_001003801
|
SMARCD3
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 3
|
|
chr16_-_31390926
|
1.196
|
|
|
|
|
chr9_+_111850770
|
1.190
|
|
AKAP2
|
A kinase (PRKA) anchor protein 2
|
|
chr21_-_26867138
|
1.186
|
NM_052954
|
CYYR1
|
cysteine/tyrosine-rich 1
|
|
chr2_+_235525355
|
1.180
|
NM_014521
|
SH3BP4
|
SH3-domain binding protein 4
|
|
chr8_-_145097031
|
1.174
|
NM_201380
|
PLEC
|
plectin
|
|
chr3_+_112273464
|
1.162
|
NM_015480
|
PVRL3
|
poliovirus receptor-related 3
|
|
chr20_+_61622985
|
1.160
|
|
PPDPF
|
pancreatic progenitor cell differentiation and proliferation factor homolog (zebrafish)
|
|
chr10_+_72102563
|
1.151
|
NM_080722
NM_139155
|
ADAMTS14
|
ADAM metallopeptidase with thrombospondin type 1 motif, 14
|
|
chr12_+_5411534
|
1.148
|
NM_001102654
|
NTF3
|
neurotrophin 3
|
|
chr9_+_112470871
|
1.141
|
NM_001166280
NM_001166281
NM_005592
|
MUSK
|
muscle, skeletal, receptor tyrosine kinase
|
|
chr7_+_114350144
|
1.138
|
|
MDFIC
|
MyoD family inhibitor domain containing
|
|
chr12_-_47869134
|
1.131
|
|
TUBA1A
|
tubulin, alpha 1a
|
|
chr11_-_110917632
|
1.129
|
|
|
|
|
chr21_-_35181308
|
1.116
|
|
RUNX1
|
runt-related transcription factor 1
|
|
chr7_+_27191534
|
1.109
|
|
HOXA11-AS1
|
HOXA11 antisense RNA 1 (non-protein coding)
|
|
chr5_-_83715947
|
1.107
|
|
EDIL3
|
EGF-like repeats and discoidin I-like domains 3
|
|
chr15_-_27901628
|
1.099
|
|
TJP1
|
tight junction protein 1 (zona occludens 1)
|
|
chr2_+_219991342
|
1.098
|
NM_001927
|
DES
|
desmin
|
|
chr7_+_27190688
|
1.085
|
|
|
|
|
chr12_-_47869067
|
1.083
|
NM_006009
|
TUBA1A
|
tubulin, alpha 1a
|
|
chr1_+_201862845
|
1.077
|
|
ATP2B4
|
ATPase, Ca++ transporting, plasma membrane 4
|
|
chr10_-_90702490
|
1.070
|
NM_001613
|
ACTA2
|
actin, alpha 2, smooth muscle, aorta
|
|
chr7_+_44110470
|
1.064
|
NM_001129
|
AEBP1
|
AE binding protein 1
|
|
chrX_+_51653348
|
1.063
|
NM_001005333
NM_006986
|
MAGED1
|
melanoma antigen family D, 1
|
|
chr19_+_16296734
|
1.062
|
|
KLF2
|
Kruppel-like factor 2 (lung)
|
|
chr9_-_112840801
|
1.052
|
|
LPAR1
|
lysophosphatidic acid receptor 1
|
|
chr21_-_35181278
|
1.047
|
|
RUNX1
|
runt-related transcription factor 1
|
|
chr7_+_116099648
|
1.041
|
NM_000245
NM_001127500
|
MET
|
met proto-oncogene (hepatocyte growth factor receptor)
|
|
chr21_-_35181094
|
1.039
|
|
|
|
|
chr11_-_11987115
|
1.037
|
|
DKK3
|
dickkopf homolog 3 (Xenopus laevis)
|
|
chrX_+_101792949
|
1.036
|
NM_001099410
NM_001099411
NM_014710
NM_001184727
|
GPRASP1
|
G protein-coupled receptor associated sorting protein 1
|
|
chrX_+_149282097
|
1.035
|
NM_001177465
|
MAMLD1
|
mastermind-like domain containing 1
|
|
chr19_+_16296632
|
1.031
|
NM_016270
|
KLF2
|
Kruppel-like factor 2 (lung)
|
|
chr16_+_31390567
|
1.027
|
NM_001042454
NM_001164719
|
TGFB1I1
|
transforming growth factor beta 1 induced transcript 1
|
|
chr1_-_39877883
|
1.024
|
|
HEYL
|
hairy/enhancer-of-split related with YRPW motif-like
|
|
chr12_+_95112385
|
1.020
|
|
ELK3
|
ELK3, ETS-domain protein (SRF accessory protein 2)
|
|
chr1_+_182622987
|
1.017
|
|
C1orf21
|
chromosome 1 open reading frame 21
|
|
chr1_-_39877928
|
1.015
|
NM_014571
|
HEYL
|
hairy/enhancer-of-split related with YRPW motif-like
|
|
chr11_-_122571046
|
1.012
|
|
CLMP
|
CXADR-like membrane protein
|
|
chr1_-_86634511
|
1.006
|
NM_001007022
NM_001184765
NM_001184766
NM_020729
|
ODF2L
|
outer dense fiber of sperm tails 2-like
|
|
chr12_+_52653176
|
1.006
|
NM_014212
|
HOXC11
|
homeobox C11
|
|
chr12_-_94466701
|
1.001
|
NM_001042403
|
USP44
|
ubiquitin specific peptidase 44
|
|
chr13_+_30378311
|
0.990
|
NM_032849
|
C13orf33
|
chromosome 13 open reading frame 33
|
|
chr12_-_50871953
|
0.990
|
NM_001081492
NM_182507
|
KRT80
|
keratin 80
|
|
chr7_-_27190882
|
0.984
|
|
HOXA11
|
homeobox A11
|
|
chr4_+_159311353
|
0.977
|
|
LOC285505
|
hypothetical protein LOC285505
|
|
chr6_+_142664490
|
0.974
|
NM_001032394
NM_001032395
NM_020455
NM_198569
|
GPR126
|
G protein-coupled receptor 126
|
|
chr3_-_107070400
|
0.974
|
NM_170662
|
CBLB
|
Cas-Br-M (murine) ecotropic retroviral transforming sequence b
|
|
chr4_+_15089931
|
0.964
|
|
CC2D2A
|
coiled-coil and C2 domain containing 2A
|
|
chr7_-_150576957
|
0.959
|
|
SMARCD3
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 3
|
|
chr1_+_201862552
|
0.953
|
|
ATP2B4
|
ATPase, Ca++ transporting, plasma membrane 4
|
|
chr4_-_149582948
|
0.951
|
NM_000901
NM_001166104
|
NR3C2
|
nuclear receptor subfamily 3, group C, member 2
|
|
chr9_+_34980266
|
0.950
|
NM_001135005
|
DNAJB5
|
DnaJ (Hsp40) homolog, subfamily B, member 5
|
|
chr6_+_17389752
|
0.946
|
NM_001143942
|
RBM24
|
RNA binding motif protein 24
|
|
chr6_+_142665057
|
0.945
|
|
GPR126
|
G protein-coupled receptor 126
|
|
chr5_+_52812165
|
0.939
|
|
FST
|
follistatin
|
|
chr2_-_175255717
|
0.936
|
NM_001077269
|
WIPF1
|
WAS/WASL interacting protein family, member 1
|
|
chrX_-_114374860
|
0.924
|
NM_020871
|
LRCH2
|
leucine-rich repeats and calponin homology (CH) domain containing 2
|
|
chr12_+_8741746
|
0.915
|
NM_020734
|
RIMKLB
|
ribosomal modification protein rimK-like family member B
|
|
chr19_-_51856129
|
0.913
|
NM_145056
|
DACT3
|
dapper, antagonist of beta-catenin, homolog 3 (Xenopus laevis)
|
|
chr2_-_175578156
|
0.913
|
NM_001025201
NM_001822
|
CHN1
|
chimerin (chimaerin) 1
|
|
chr13_-_109236897
|
0.904
|
NM_003749
|
IRS2
|
insulin receptor substrate 2
|
|
chr7_-_27120015
|
0.898
|
|
HOXA3
|
homeobox A3
|
|
chrX_+_133993984
|
0.887
|
NM_001078171
|
FAM127A
|
family with sequence similarity 127, member A
|
|
chr10_+_52420936
|
0.885
|
NM_001098512
|
PRKG1
|
protein kinase, cGMP-dependent, type I
|
|
chr6_-_85529884
|
0.883
|
|
TBX18
|
T-box 18
|
|
chr11_-_27697768
|
0.880
|
NM_001143807
|
BDNF
|
brain-derived neurotrophic factor
|
|
chr1_+_199519202
|
0.876
|
NM_000299
NM_001005337
|
PKP1
|
plakophilin 1 (ectodermal dysplasia/skin fragility syndrome)
|
|
chr5_-_156935283
|
0.876
|
|
|
|
|
chr5_+_140835788
|
0.874
|
|
PCDHGC3
|
protocadherin gamma subfamily C, 3
|
|
chr1_+_181259317
|
0.866
|
|
|
|
|
chr4_-_149582754
|
0.864
|
|
NR3C2
|
nuclear receptor subfamily 3, group C, member 2
|
|
chr17_+_52026058
|
0.863
|
NM_005450
|
NOG
|
noggin
|
|
chrX_-_48818585
|
0.862
|
NM_007213
|
PRAF2
|
PRA1 domain family, member 2
|
|
chr1_+_9275542
|
0.862
|
|
SPSB1
|
splA/ryanodine receptor domain and SOCS box containing 1
|
|
chrX_+_101887562
|
0.861
|
NM_001142529
NM_001142530
|
BHLHB9
|
basic helix-loop-helix domain containing, class B, 9
|
|
chr12_-_48902692
|
0.859
|
NM_001113547
|
LIMA1
|
LIM domain and actin binding 1
|
|
chr12_-_21985433
|
0.859
|
|
ABCC9
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 9
|
|
chr4_+_123967389
|
0.856
|
|
FGF2
|
fibroblast growth factor 2 (basic)
|
|
chr9_+_101623957
|
0.856
|
NM_006981
NM_173199
|
NR4A3
|
nuclear receptor subfamily 4, group A, member 3
|
|
chr5_+_131621276
|
0.852
|
|
PDLIM4
|
PDZ and LIM domain 4
|
|
chr7_+_79602023
|
0.847
|
NM_002069
|
GNAI1
|
guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 1
|
|
chr17_+_56832492
|
0.841
|
|
TBX2
|
T-box 2
|
|
chr9_-_122516399
|
0.838
|
NM_001080497
|
MEGF9
|
multiple EGF-like-domains 9
|
|
chr7_-_27102049
|
0.838
|
NM_005522
NM_153620
|
HOXA1
|
homeobox A1
|
|
chr8_-_134653194
|
0.836
|
NM_003033
NM_173344
|
ST3GAL1
|
ST3 beta-galactoside alpha-2,3-sialyltransferase 1
|
|
chr17_+_56884560
|
0.834
|
|
TBX4
|
T-box 4
|
|
chr7_-_92301147
|
0.830
|
NM_001259
|
CDK6
|
cyclin-dependent kinase 6
|
|
chr14_+_105012106
|
0.828
|
NM_001312
|
CRIP2
|
cysteine-rich protein 2
|
|
chrX_-_134013644
|
0.826
|
NM_001078172
NM_001134321
|
FAM127B
|
family with sequence similarity 127, member B
|
|
chr13_+_75108450
|
0.825
|
|
LMO7
|
LIM domain 7
|
|
chr19_-_54557450
|
0.825
|
NM_003598
|
TEAD2
|
TEA domain family member 2
|
|
chrX_-_133984157
|
0.824
|
NM_001078173
|
FAM127C
|
family with sequence similarity 127, member C
|
|
chr13_+_41520888
|
0.807
|
NM_152910
NM_178009
|
DGKH
|
diacylglycerol kinase, eta
|
|
chr1_+_9275518
|
0.806
|
NM_025106
|
SPSB1
|
splA/ryanodine receptor domain and SOCS box containing 1
|
|
chr1_+_17817397
|
0.806
|
|
ARHGEF10L
|
Rho guanine nucleotide exchange factor (GEF) 10-like
|
|
chr11_-_44928803
|
0.804
|
NM_001076787
NM_006034
|
TP53I11
|
tumor protein p53 inducible protein 11
|
|
chr12_+_54400417
|
0.804
|
NM_002905
|
RDH5
|
retinol dehydrogenase 5 (11-cis/9-cis)
|
|
chr5_+_131621230
|
0.803
|
NM_001131027
NM_003687
|
PDLIM4
|
PDZ and LIM domain 4
|
|
chr11_-_11987198
|
0.802
|
NM_015881
|
DKK3
|
dickkopf homolog 3 (Xenopus laevis)
|
|
chr10_+_75343352
|
0.801
|
|
PLAU
|
plasminogen activator, urokinase
|
|
chr2_-_133144094
|
0.798
|
|
LYPD1
|
LY6/PLAUR domain containing 1
|
|
chr5_+_140835727
|
0.797
|
NM_002588
NM_032402
NM_032403
|
PCDHGC3
|
protocadherin gamma subfamily C, 3
|
|
chr3_-_113841200
|
0.794
|
|
CCDC80
|
coiled-coil domain containing 80
|
|
chr19_+_10597167
|
0.793
|
NM_020428
|
SLC44A2
|
solute carrier family 44, member 2
|
|
chr17_-_72045218
|
0.791
|
NM_134268
|
CYGB
|
cytoglobin
|
|
chr17_+_35751796
|
0.788
|
NM_001024809
|
RARA
|
retinoic acid receptor, alpha
|
|
chr12_+_95112131
|
0.788
|
NM_005230
|
ELK3
|
ELK3, ETS-domain protein (SRF accessory protein 2)
|
|
chr2_+_202607554
|
0.786
|
NM_003507
|
FZD7
|
frizzled homolog 7 (Drosophila)
|
|
chr1_-_204048758
|
0.785
|
NM_173854
|
SLC41A1
|
solute carrier family 41, member 1
|
|
chr11_-_66898223
|
0.782
|
NM_001166212
|
CLCF1
|
cardiotrophin-like cytokine factor 1
|
|
chrX_+_101853912
|
0.782
|
NM_001004051
NM_001184874
NM_001184875
NM_001184876
NM_138437
|
GPRASP2
|
G protein-coupled receptor associated sorting protein 2
|
|
chr6_+_1335067
|
0.780
|
NM_001452
|
FOXF2
|
forkhead box F2
|
|
chr6_+_133604181
|
0.779
|
NM_004100
NM_172103
NM_172105
|
EYA4
|
eyes absent homolog 4 (Drosophila)
|
|
chr4_+_123967265
|
0.775
|
NM_002006
|
FGF2
|
fibroblast growth factor 2 (basic)
|
|
chr6_-_112682401
|
0.771
|
|
LAMA4
|
laminin, alpha 4
|
|
chr17_-_15185623
|
0.768
|
NM_031898
|
TEKT3
|
tektin 3
|
|
chr5_+_131621287
|
0.764
|
|
PDLIM4
|
PDZ and LIM domain 4
|
|
chr1_-_93919789
|
0.764
|
NM_003567
|
BCAR3
|
breast cancer anti-estrogen resistance 3
|
|
chr2_-_9061193
|
0.763
|
NM_138799
|
MBOAT2
|
membrane bound O-acyltransferase domain containing 2
|
|
chr3_-_180652024
|
0.762
|
NM_021629
|
GNB4
|
guanine nucleotide binding protein (G protein), beta polypeptide 4
|
|
chr19_-_60573552
|
0.758
|
NM_000641
|
IL11
|
interleukin 11
|
|
chr8_-_13035102
|
0.756
|
NM_006094
|
DLC1
|
deleted in liver cancer 1
|
|
chr14_+_104218487
|
0.754
|
|
|
|
|
chr1_-_29321599
|
0.753
|
NM_001171868
|
TMEM200B
|
transmembrane protein 200B
|
|
chr19_-_13910167
|
0.751
|
NM_024825
|
PODNL1
|
podocan-like 1
|
|
chr1_+_155130146
|
0.750
|
NM_001080471
|
PEAR1
|
platelet endothelial aggregation receptor 1
|
|
chr1_+_234372454
|
0.730
|
NM_003272
|
GPR137B
|
G protein-coupled receptor 137B
|
|
chr6_-_112682459
|
0.730
|
NM_001105206
NM_001105207
NM_001105208
NM_001105209
NM_002290
|
LAMA4
|
laminin, alpha 4
|
|
chr11_-_61415429
|
0.728
|
NM_021727
|
FADS3
|
fatty acid desaturase 3
|
|
chr3_-_180651874
|
0.725
|
|
GNB4
|
guanine nucleotide binding protein (G protein), beta polypeptide 4
|
|
chr8_-_119703297
|
0.724
|
|
SAMD12
|
sterile alpha motif domain containing 12
|
|
chr19_-_13808101
|
0.724
|
|
LOC284454
|
hypothetical LOC284454
|
|
chr5_+_52812236
|
0.721
|
NM_006350
NM_013409
|
FST
|
follistatin
|
|
chr8_+_70541631
|
0.721
|
|
SULF1
|
sulfatase 1
|
|
chr12_+_12935619
|
0.720
|
|
GPRC5A
|
G protein-coupled receptor, family C, group 5, member A
|