Mean activities
Top 10 and bottom 10 mean activities.
Mean activities sorted by motif Z-value.
| Motif name | Mean activity | Z-Value | Associated genes | Logo |
|---|---|---|---|---|
| AAAGUGC | -0.047 | -0.131 |
|

|
| AACACUG | -0.322 | -0.624 |
|

|
| AACAGUC | -0.056 | -0.067 |
|

|
| AACCGUU | -2.055 | -0.271 |
|
|
| AACCUGG | 0.772 | 0.257 |
|
|
| AACGGAA | 0.655 | 0.249 |
|
|
| AAGGCAC | 0.057 | 0.193 |
|
|
| AAGGUGC | 0.233 | 0.282 |
|
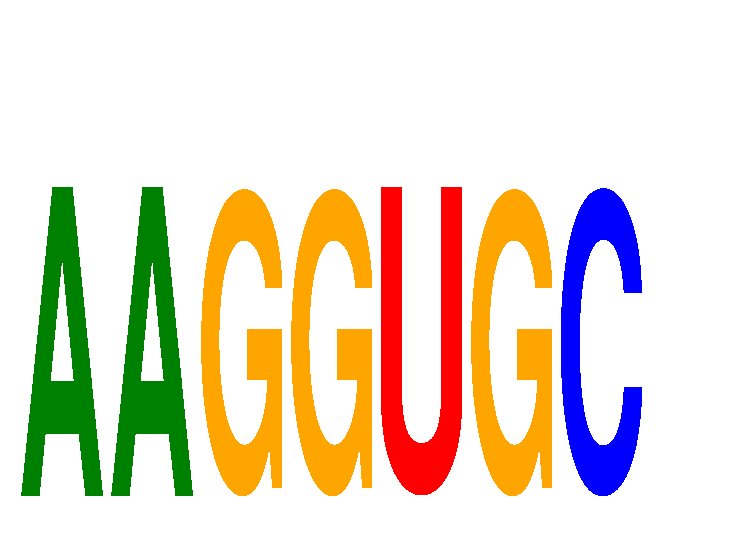
|
| AAGUGCU | -0.430 | -0.873 |
|

|
| AAUACUG | 0.229 | 0.652 |
|

|
| AAUCUCA | -0.304 | -0.183 |
|

|
| AAUCUCU | 0.359 | 0.195 |
|

|
| AAUGCCC | -0.065 | -0.070 |
|
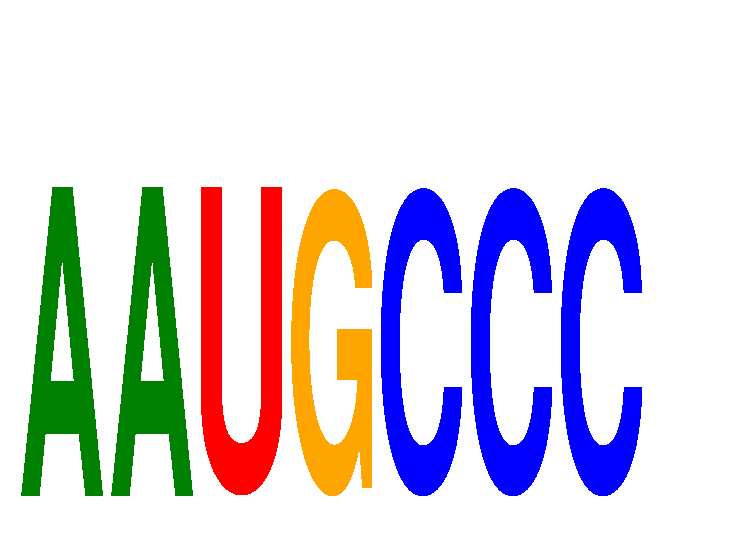
|
| ACAGUAC | -0.147 | -0.268 |
|

|
| ACAGUAU | -0.081 | -0.143 |
|

|
| ACAUUCA | 0.078 | 0.206 |
|

|
| ACCACAG | 0.037 | 0.026 |
|

|
| ACCCGUA | -1.737 | -0.326 |
|
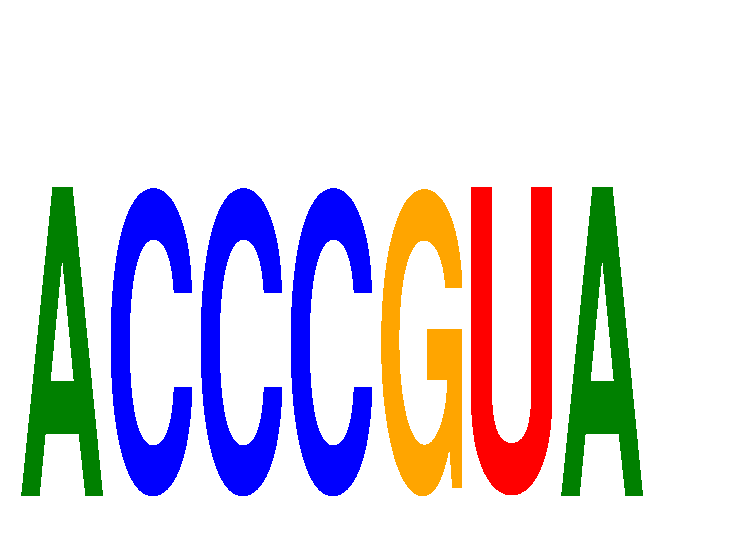
|
| ACCCUGU | -0.133 | -0.148 |
|
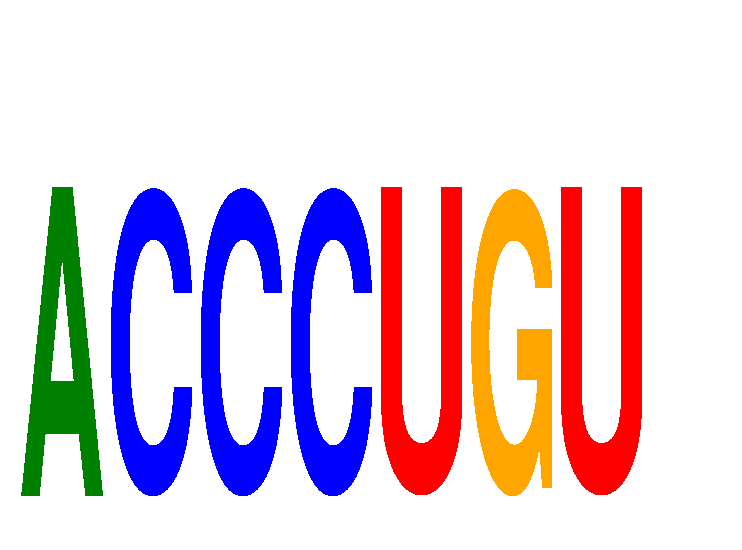
|
| ACUGCAU | 0.588 | 0.535 |
|

|
| ACUGGCC | -0.404 | -0.394 |
|

|
| AGAUCAG | 0.856 | 0.128 |
|

|
| AGCACCA | -0.037 | -0.125 |
|

|
| AGCAGCA | 0.065 | 0.165 |
|

|
| AGCAGCG | -0.278 | -0.463 |
|

|
| AGCCCUU | -0.335 | -0.549 |
|

|
| AGCUGCC | -0.247 | -0.462 |
|

|
| AGCUUAU | -0.025 | -0.029 |
|

|
| AGGUAGU | -0.222 | -0.344 |
|

|
| AGUGCAA | -0.180 | -0.451 |
|

|
| AGUGCUU | 0.248 | 0.485 |
|

|
| AGUGGUU | -0.273 | -0.390 |
|

|
| AHR_ARNT2 | 0.011 | 0.108 |
|
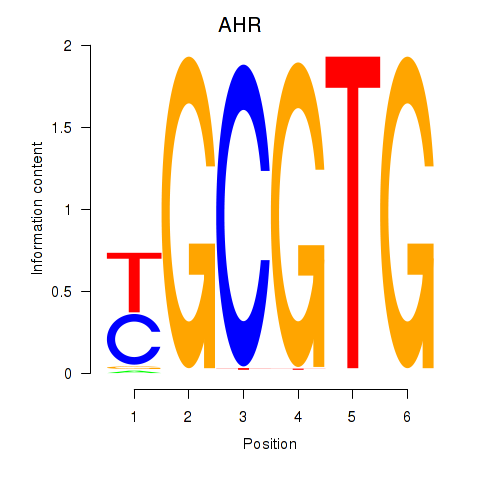
|
| AIRE | -0.157 | -0.269 |
|
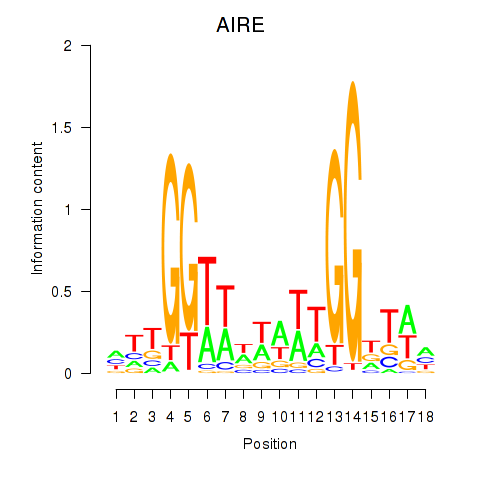
|
| ALX1_ARX | -0.124 | -0.135 |
|
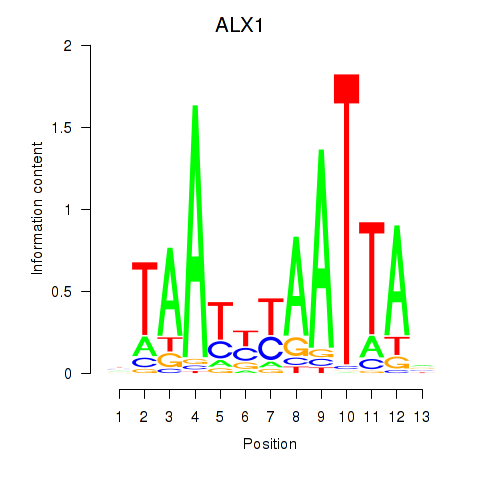
|
| ALX3 | -0.069 | -0.122 |
|
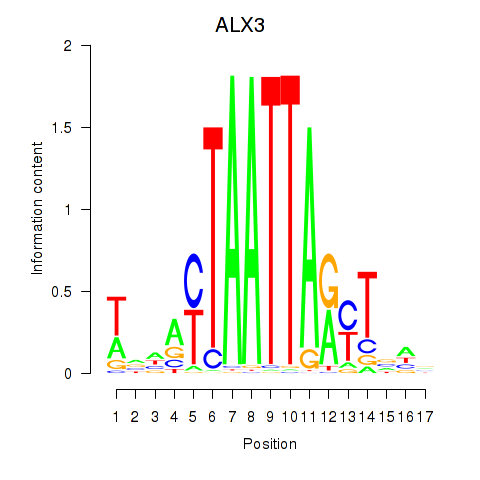
|
| ARID3A | -0.017 | -0.053 |
|
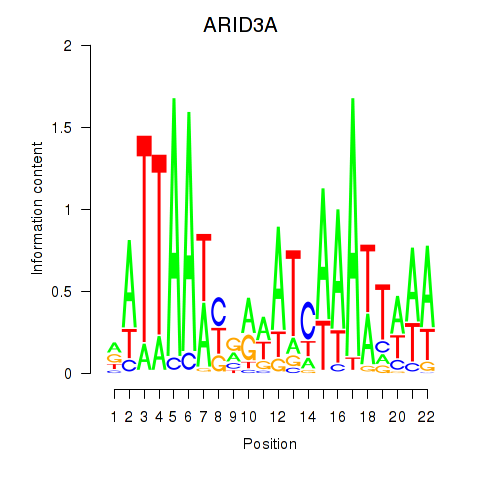
|
| ARID5A | -0.040 | -0.128 |
|

|
| ARID5B | 0.239 | 0.484 |
|
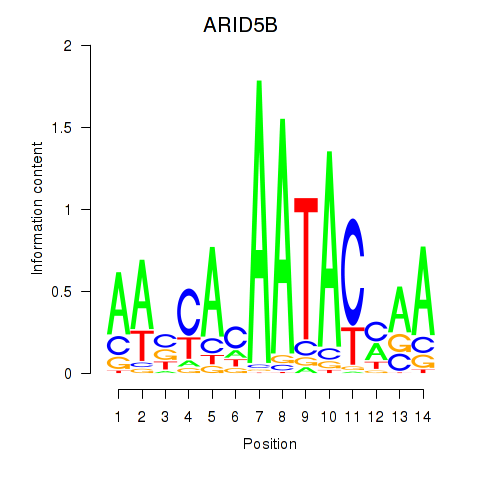
|
| ARNT | 0.408 | 1.311 |
|
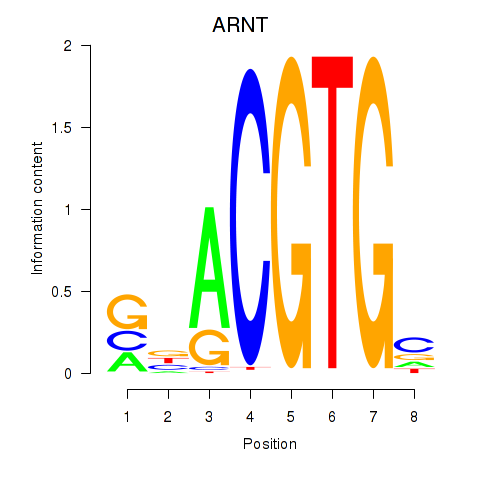
|
| AR_NR3C2 | -0.098 | -0.171 |
|
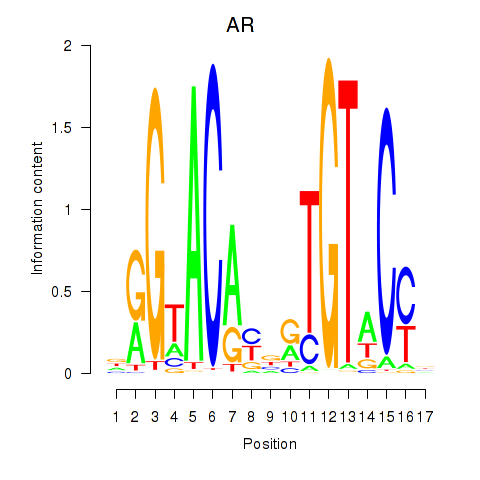
|
| ATF2_ATF1_ATF3 | 0.139 | 0.865 |
|
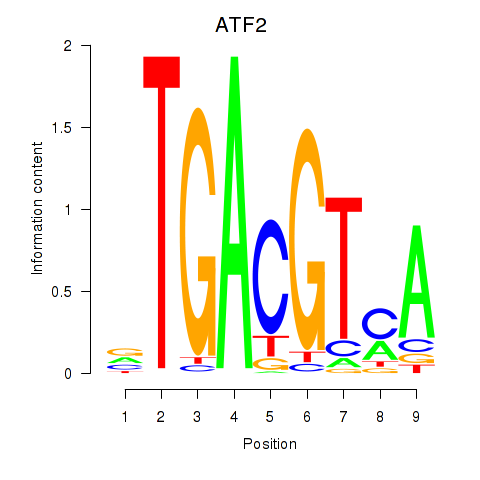
|
| ATF4 | 0.524 | 0.823 |
|
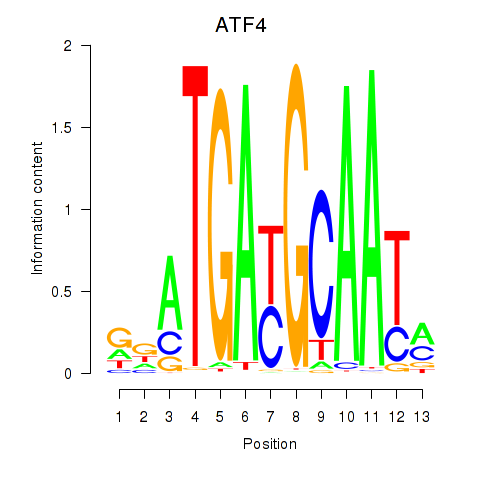
|
| ATF5 | 0.028 | 0.061 |
|
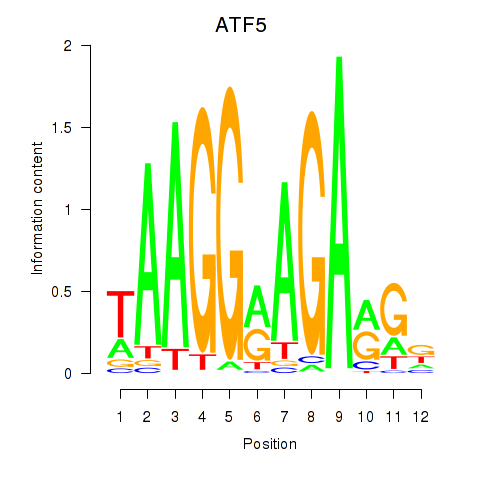
|
| ATF6 | 0.270 | 0.868 |
|
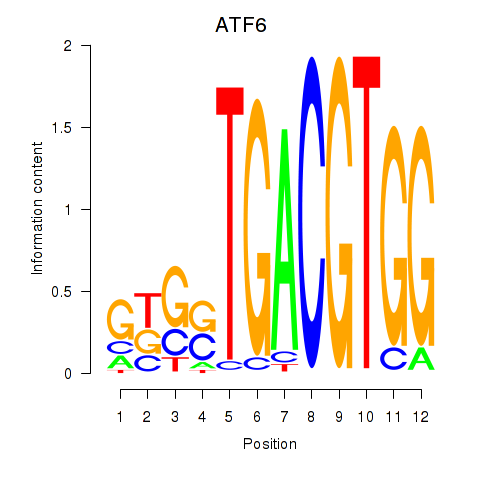
|
| ATF7 | -0.228 | -0.521 |
|
|
| AUAAAGU | 0.654 | 1.010 |
|

|
| AUGACAC | 2.018 | 0.489 |
|

|
| AUGGCAC | 0.847 | 1.337 |
|

|
| AUGGCUU | -0.285 | -0.704 |
|

|
| AUGUGCC | 0.410 | 0.347 |
|

|
| AUUGCAC | 0.054 | 0.149 |
|

|
| BACH1_NFE2_NFE2L2 | 0.726 | 2.079 |
|
|
| BACH2 | 0.433 | 0.873 |
|
|
| BARHL1 | 0.149 | 1.189 |
|
|
| BARHL2 | -0.099 | -0.339 |
|
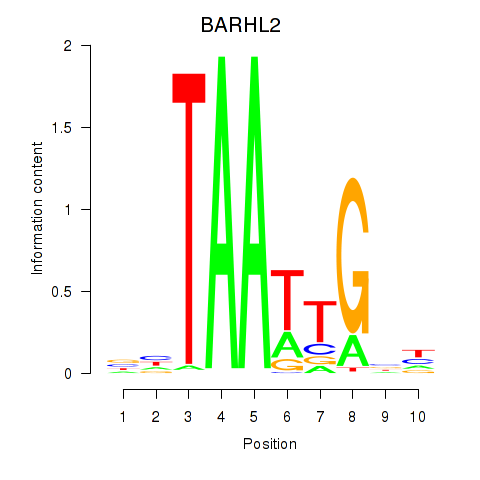
|
| BARX1 | 0.193 | 0.791 |
|
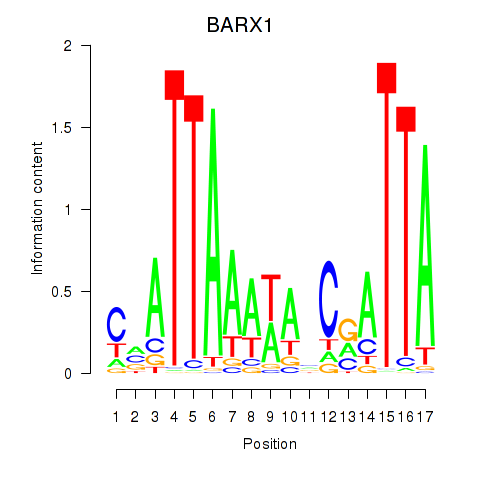
|
| BATF | 0.187 | 1.280 |
|
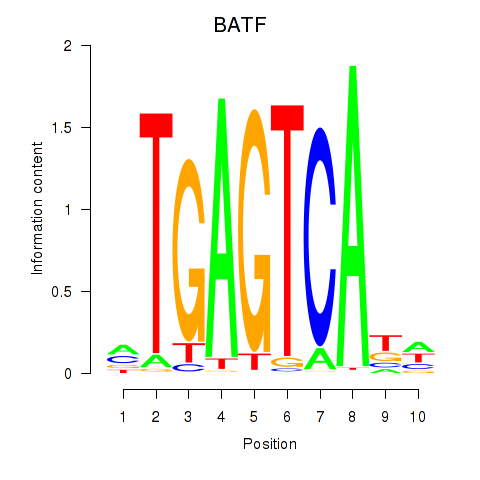
|
| BATF3 | 0.157 | 0.189 |
|

|
| BBX | 0.362 | 0.492 |
|
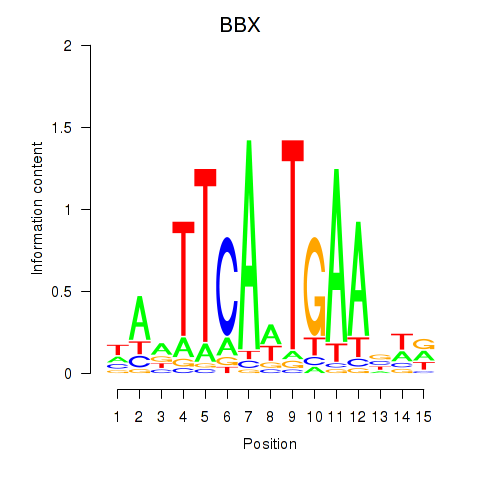
|
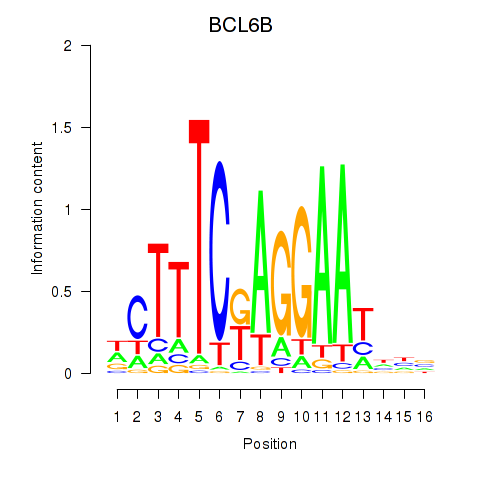
| BCL6B | 0.593 | 1.088 |
|
|
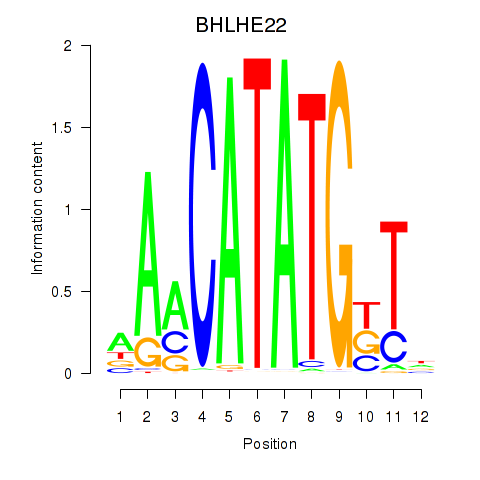
| BHLHE22_BHLHA15_BHLHE23 | -0.138 | -0.201 |
|
|
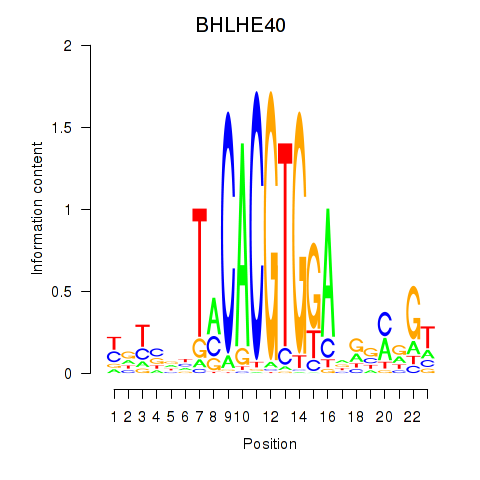
| BHLHE40 | 0.188 | 0.350 |
|
|
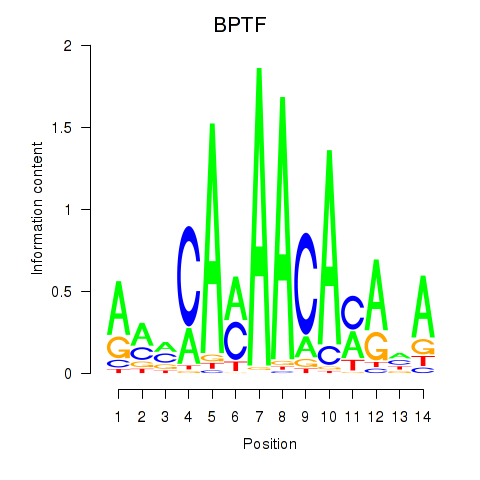
| BPTF | 0.113 | 0.939 |
|
|
| BRCA1 | -0.128 | -0.157 |
|
|
| CACAGUG | 0.262 | 0.486 |
|

|
| CACCUCC | 4.240 | 0.932 |
|

|
| CAGUAGU | -0.186 | -0.326 |
|

|
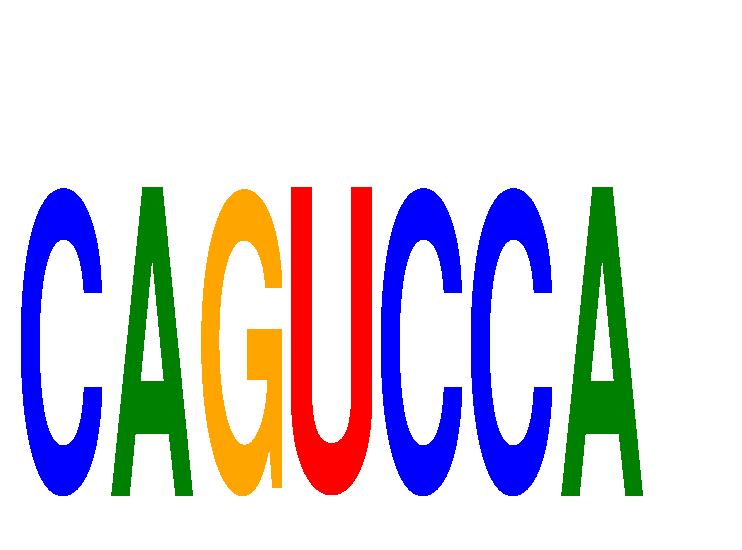
| CAGUCCA | 0.093 | 0.076 |
|

|
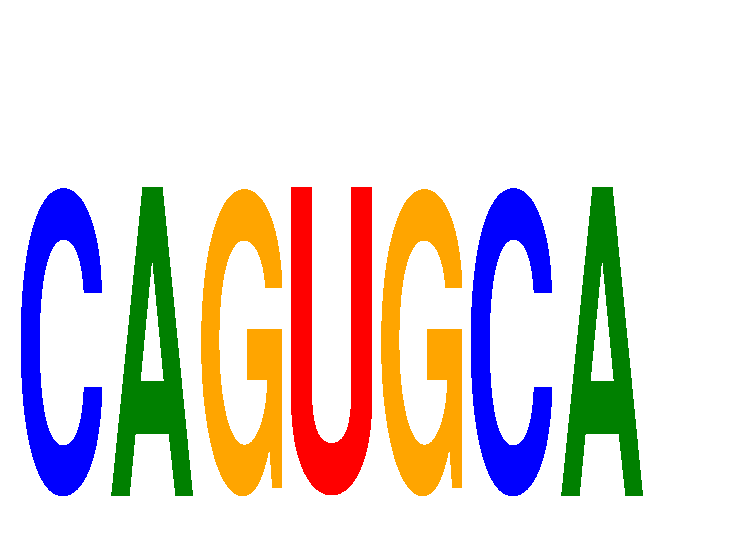
| CAGUGCA | 0.022 | 0.049 |
|

|
| CBFB | 0.215 | 0.256 |
|
|
| CCACAGG | -0.385 | -0.267 |
|

|
| CCAGCAU | -0.005 | -0.003 |
|

|
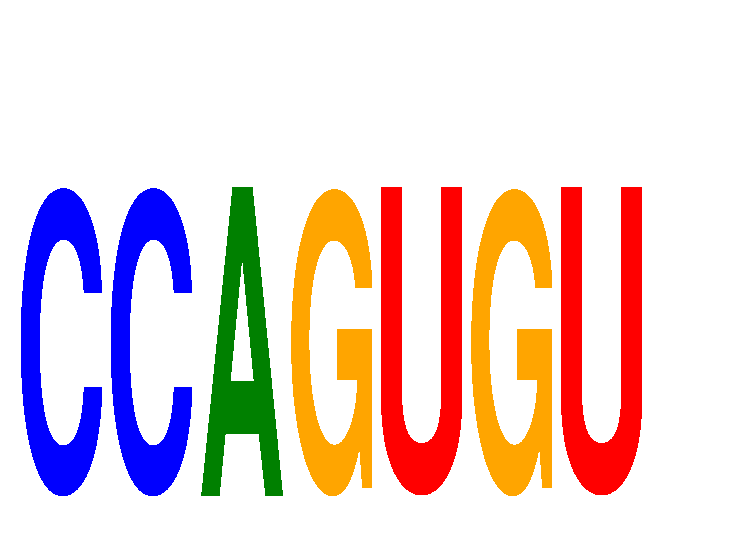
| CCAGUGU | -0.097 | -0.190 |
|

|
| CCCUGAG | -0.581 | -1.534 |
|
|
| CCUUCAU | 0.614 | 0.634 |
|
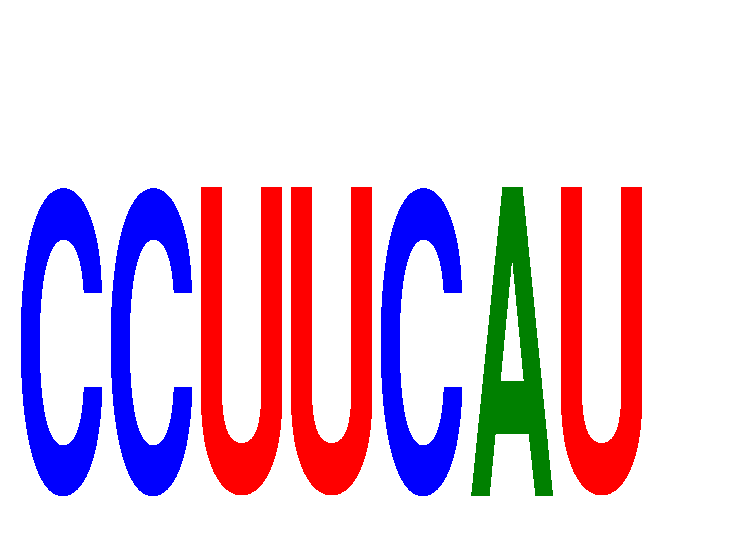

|
| CCUUGGC | -0.604 | -0.227 |
|

|
| CDC5L | -0.178 | -0.668 |
|
|
| CDX1 | 0.013 | 0.038 |
|
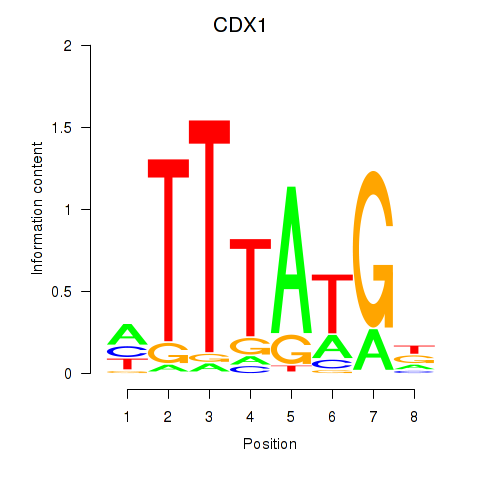
|
| CDX2 | 0.291 | 0.519 |
|
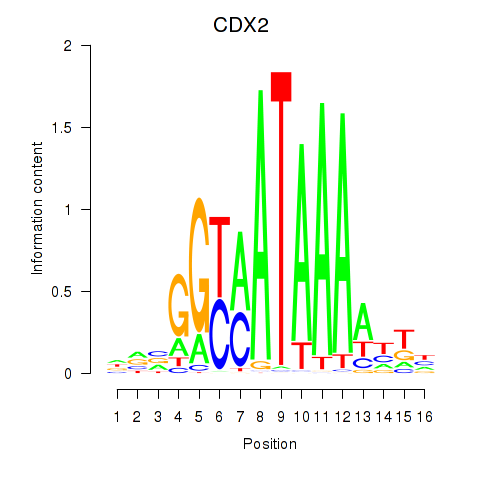
|
| CEBPA | -0.195 | -0.666 |
|
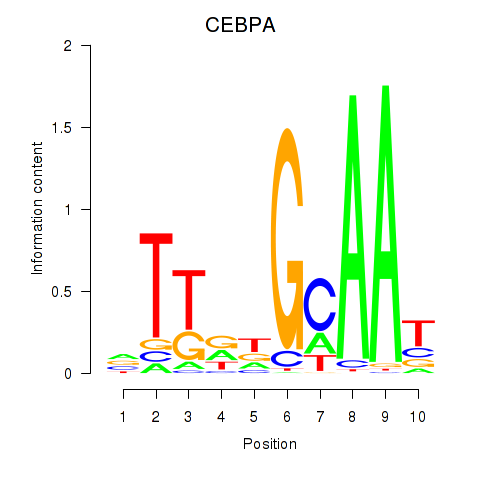
|
| CEBPB | 0.040 | 0.088 |
|
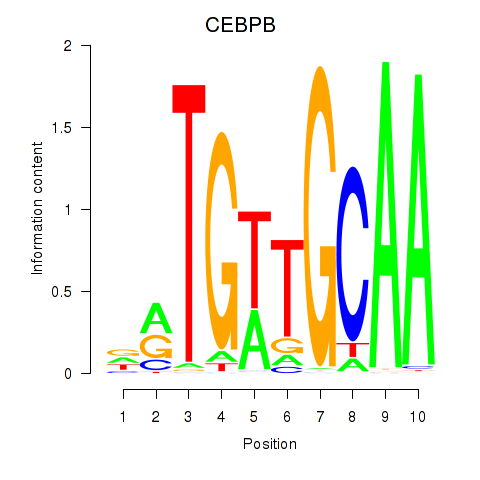
|
| CEBPE_CEBPD | 0.052 | 0.145 |
|
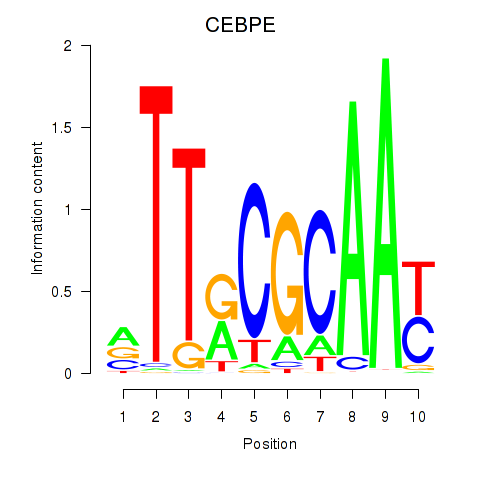
|
| CEBPG | -0.046 | -0.081 |
|
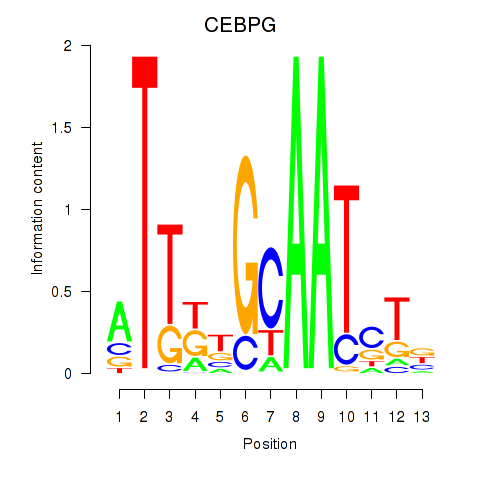
|
| CENPB | -0.102 | -0.185 |
|
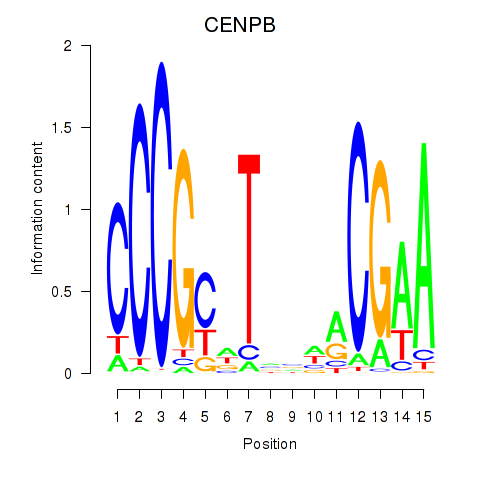
|
| CGUACCG | -0.645 | -0.179 |
|

|
| CGUGUCU | -0.534 | -0.069 |
|

|
| CLOCK | 0.162 | 0.542 |
|
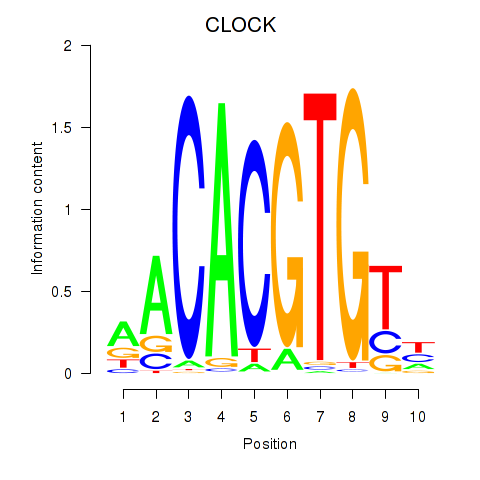
|
| CPEB1 | 0.019 | 0.070 |
|
|
| CREB1 | 0.137 | 0.440 |
|
|
| CREB3L1_CREB3 | 0.377 | 0.966 |
|
|
| CREB3L2 | 0.162 | 0.432 |
|
|
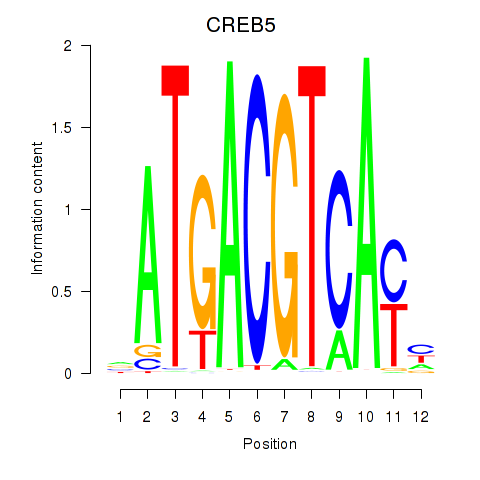
| CREB5_CREM_JUNB | -0.052 | -0.178 |
|
|
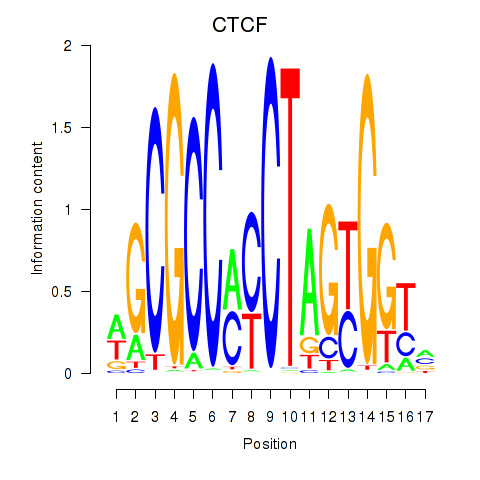
| CTCF_CTCFL | -0.005 | -0.020 |
|
|
| CUACAGU | -0.264 | -0.116 |
|

|
| CUGGCUG | -0.407 | -0.371 |
|

|
| CUUUGGU | -0.119 | -0.393 |
|

|
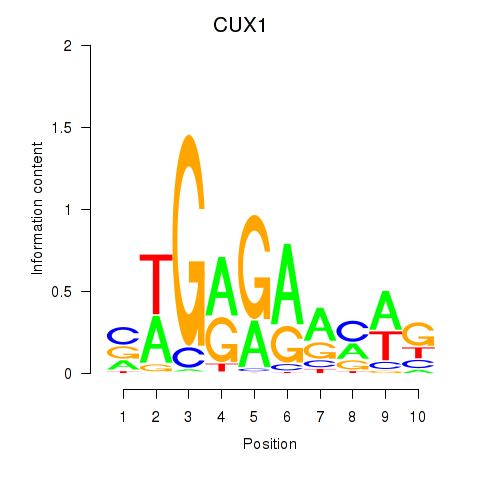
| CUX1 | -0.222 | -0.920 |
|
|
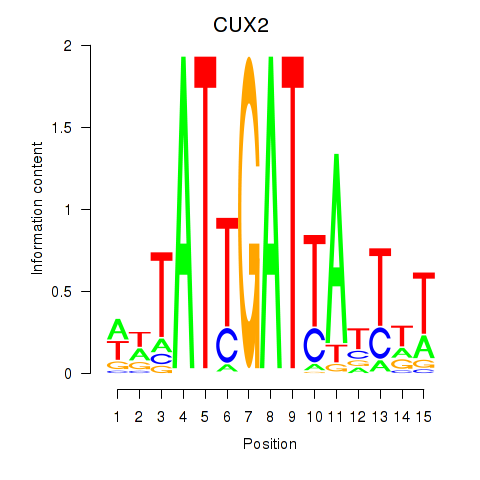
| CUX2 | -0.229 | -0.875 |
|
|
| CXXC1 | 0.293 | 1.681 |
|
|
| DBP | 0.006 | 0.017 |
|
|
| DBX2_HLX | 0.063 | 0.165 |
|
|
| DDIT3 | 0.323 | 0.479 |
|
|
| DLX1_HOXA3_BARX2 | -0.071 | -0.177 |
|
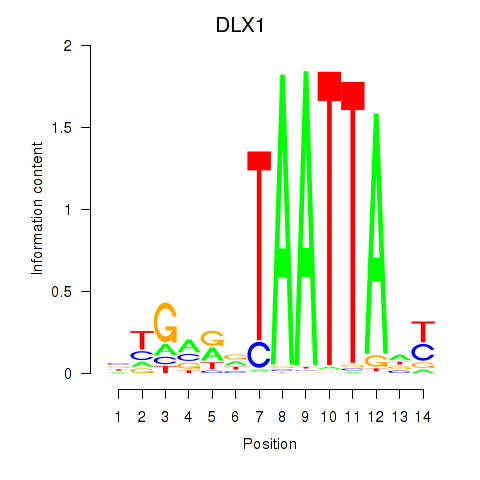
|
| DLX3_EVX1_MEOX1 | 0.390 | 0.609 |
|
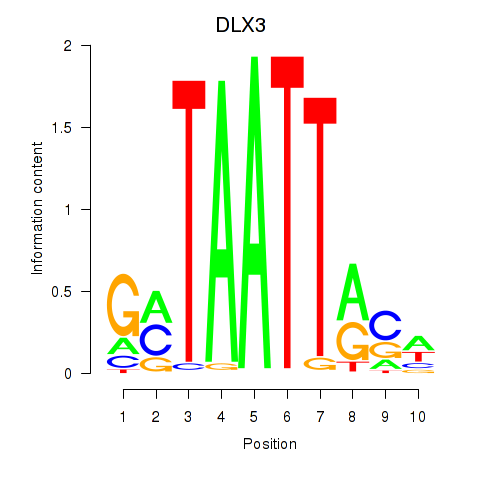
|
| DLX4_HOXD8 | -0.186 | -0.900 |
|
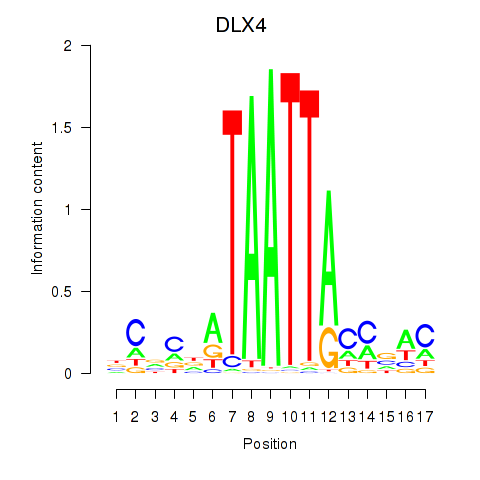
|
| DLX5 | -0.075 | -0.093 |
|
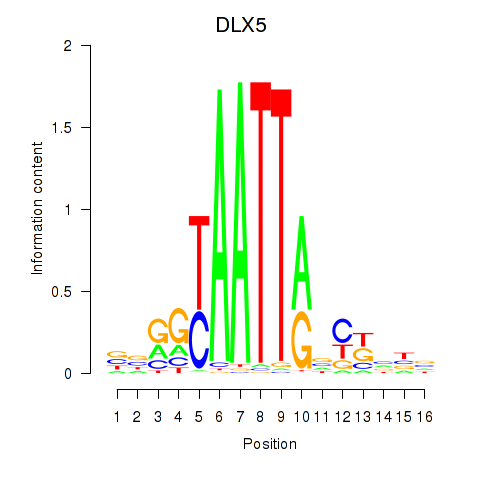
|
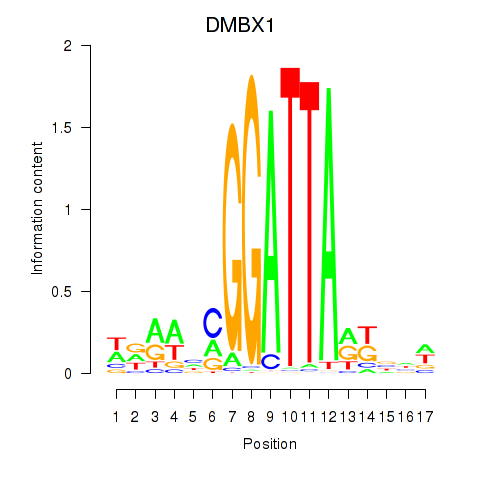
| DMBX1 | 0.133 | 0.155 |
|
|
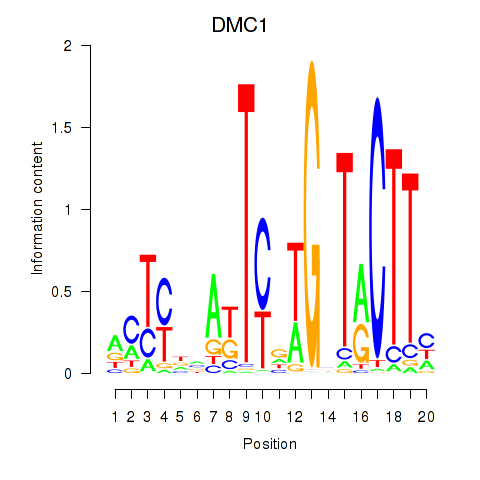
| DMC1 | 0.401 | 0.516 |
|
|
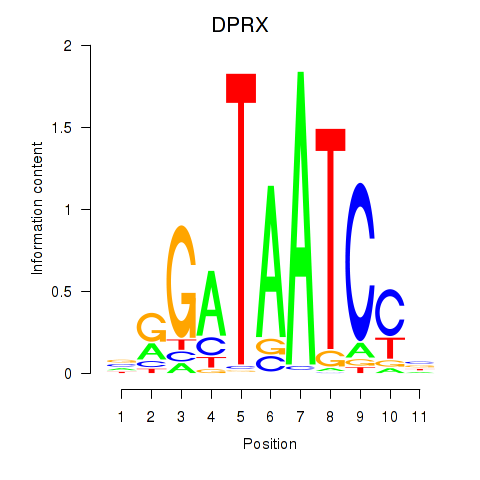
| DPRX | 0.223 | 0.651 |
|
|
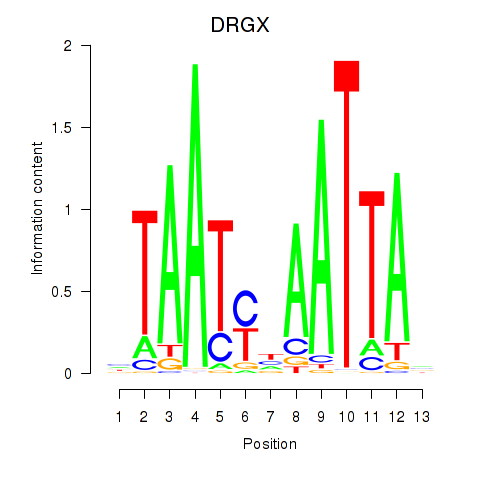
| DRGX_PROP1 | -0.794 | -1.124 |
|
|
| DUXA | -0.363 | -1.165 |
|
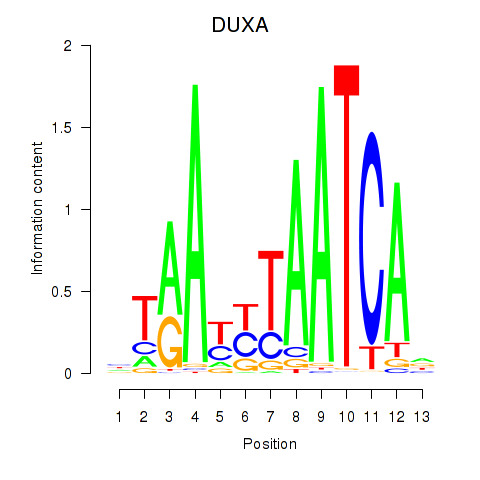
|
| E2F2_E2F5 | 0.109 | 0.356 |
|
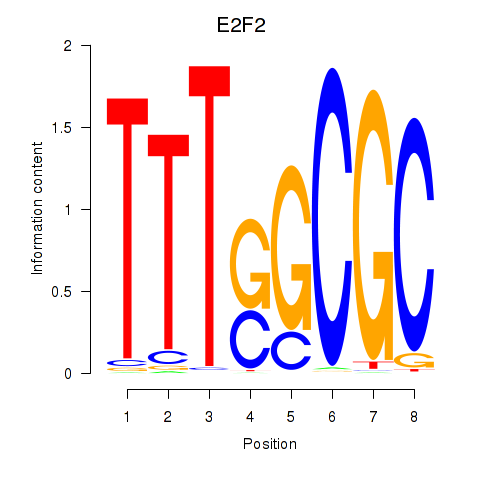
|
| E2F3 | 0.191 | 1.574 |
|
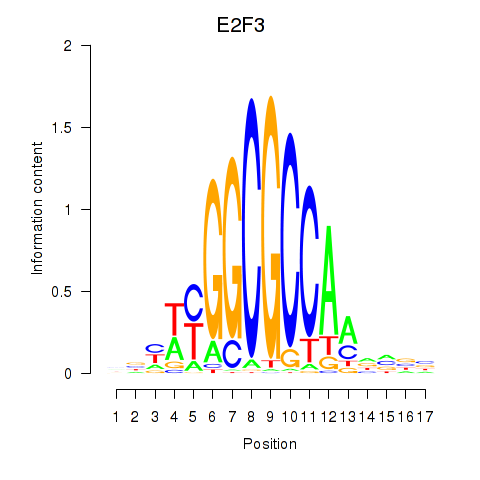
|
| E2F4 | -0.068 | -0.217 |
|
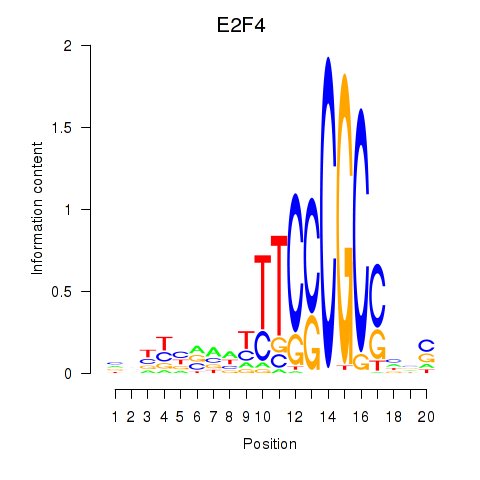
|
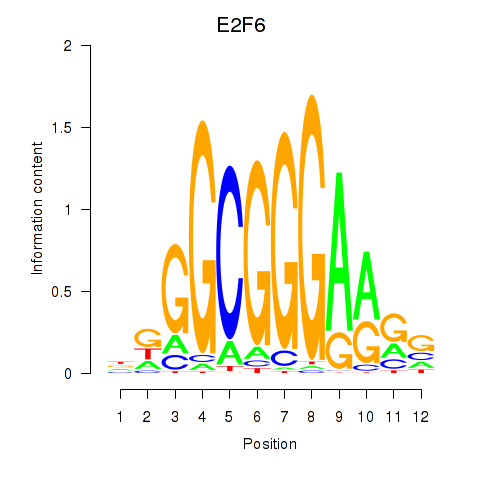
| E2F6 | -0.052 | -0.435 |
|
|
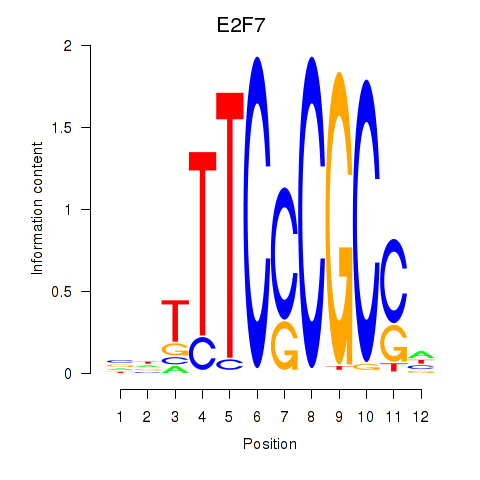
| E2F7_E2F1 | 0.107 | 0.716 |
|
|
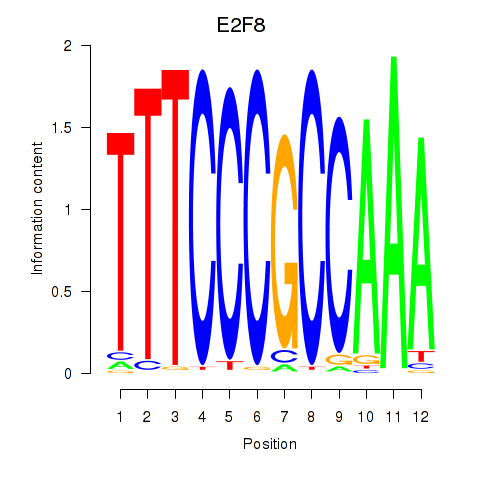
| E2F8 | 0.251 | 0.790 |
|
|
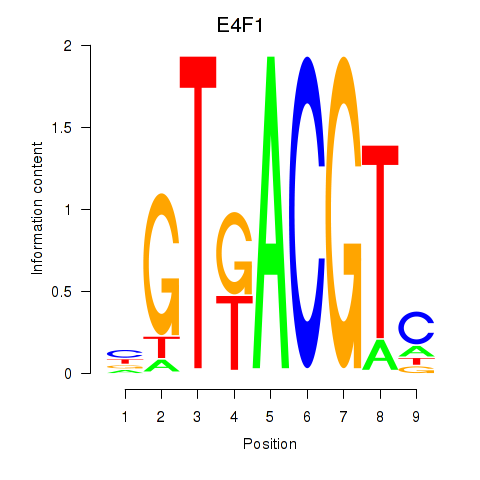
| E4F1 | 0.024 | 0.036 |
|
|
| EBF1 | -0.209 | -1.711 |
|
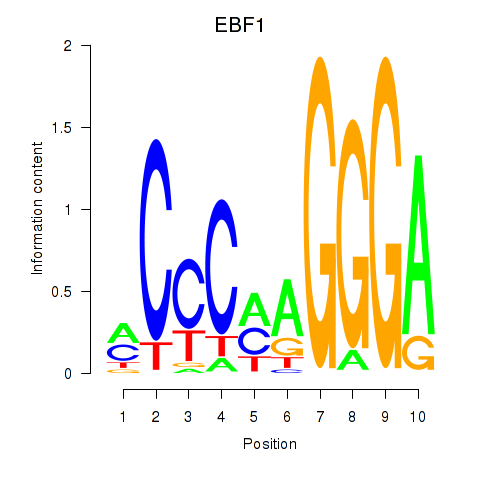
|
| EBF3 | 0.073 | 0.269 |
|
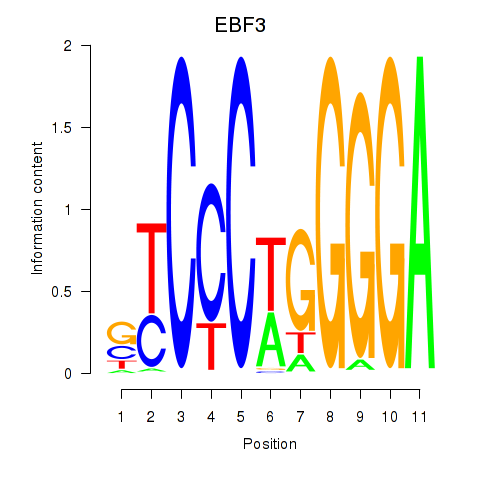
|
| EGR1_EGR4 | -0.072 | -0.281 |
|
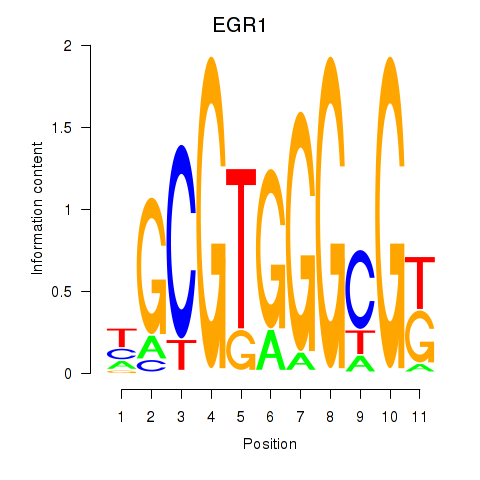
|
| EGR3_EGR2 | 0.135 | 0.508 |
|
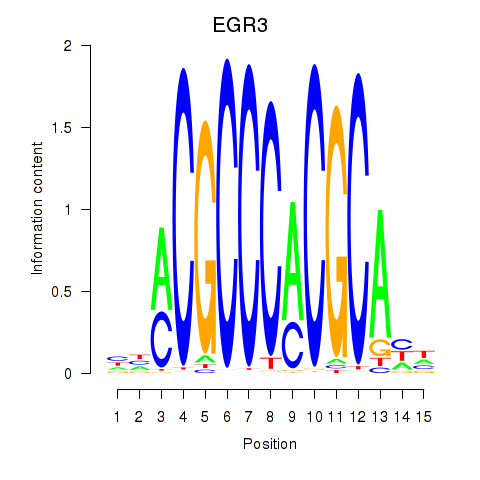
|
| ELF2_GABPA_ELF5 | 0.156 | 1.624 |
|
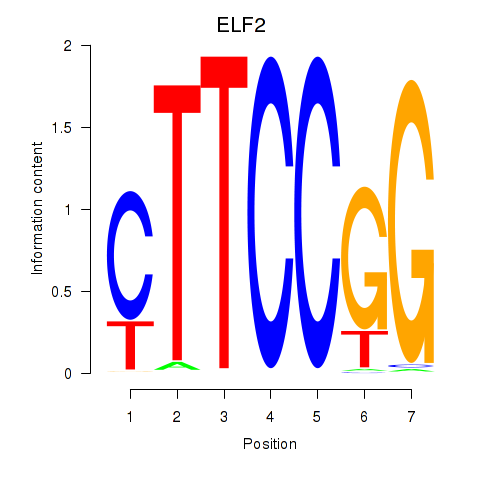
|
| ELF3_EHF | -0.086 | -0.458 |
|
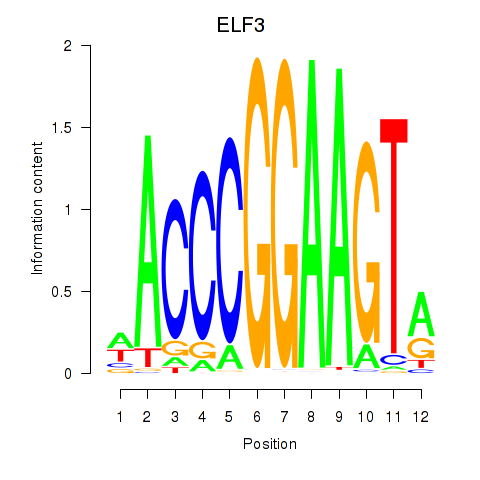
|
| ELK4_ETV5_ELK1_ELK3_ELF4 | 0.275 | 2.340 |
|

|
| EMX1 | -0.187 | -0.430 |
|
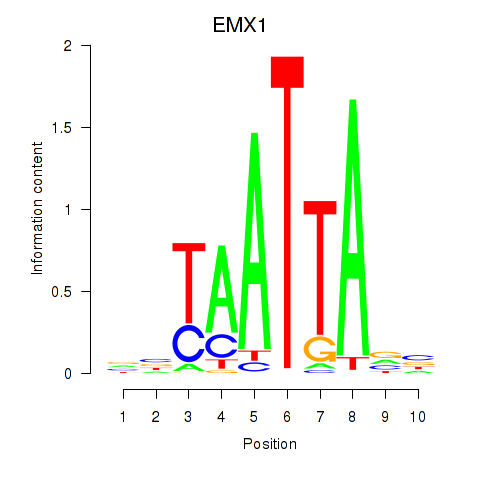
|
| EMX2 | -0.143 | -0.407 |
|
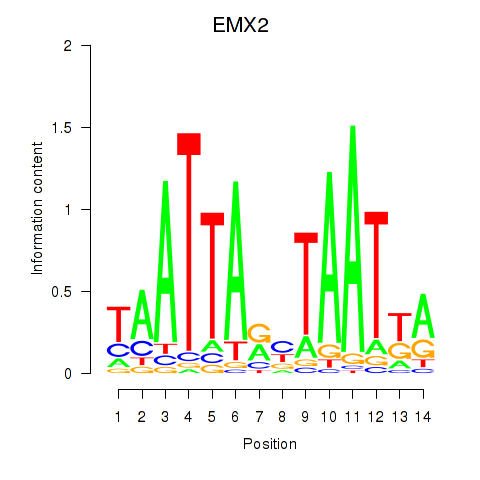
|
| EN1_ESX1_GBX1 | -0.270 | -0.585 |
|
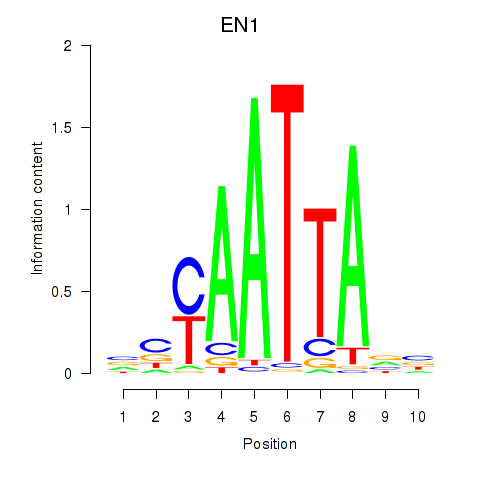
|
| EN2_GBX2_LBX2 | 0.081 | 0.119 |
|
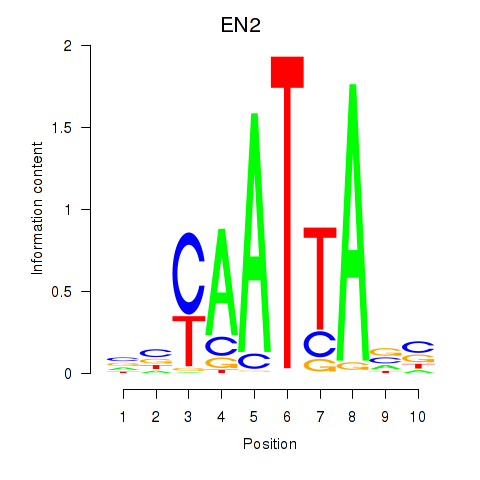
|
| EOMES | 0.157 | 0.601 |
|
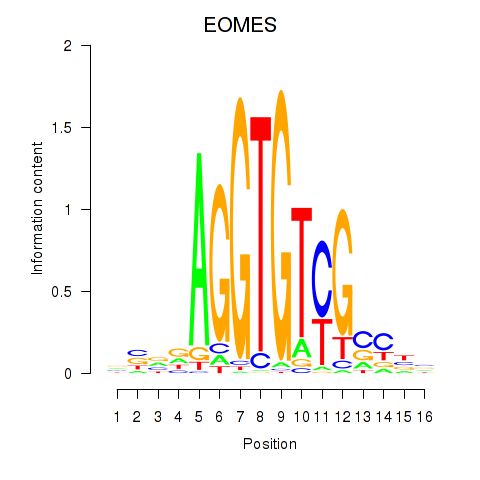
|
| EP300 | -0.001 | -0.004 |
|
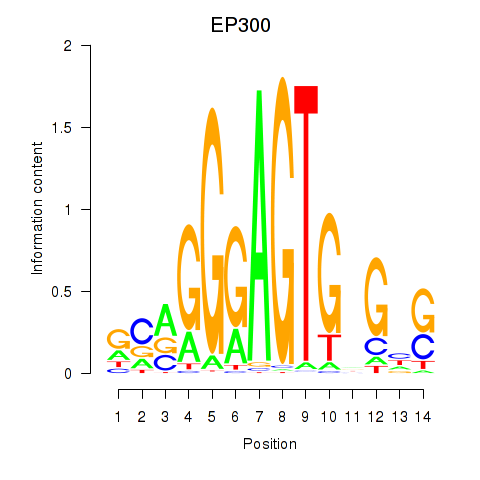
|
| EPAS1_BCL3 | -0.404 | -2.539 |
|
|
| ERG | -0.097 | -0.696 |
|
|
| ESR1 | -0.075 | -0.328 |
|
|
| ESRRA_ESR2 | -0.111 | -1.454 |
|
|
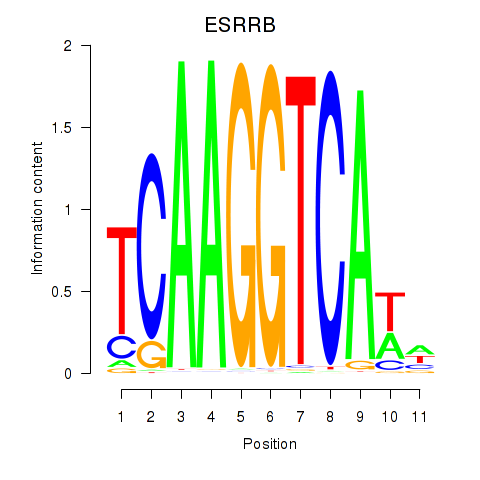
| ESRRB_ESRRG | 0.225 | 0.801 |
|
|
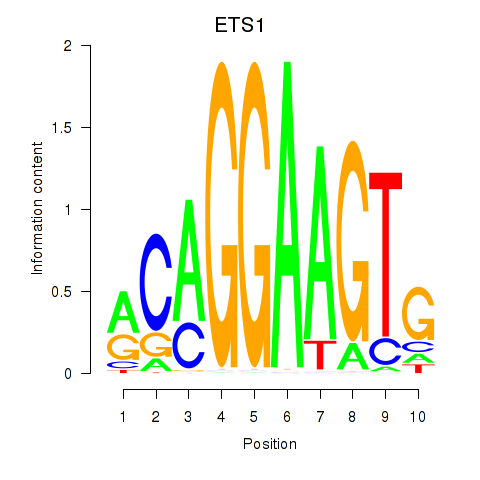
| ETS1 | 0.185 | 0.943 |
|
|
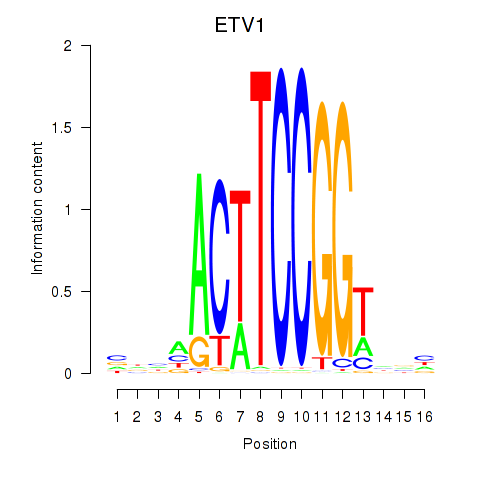
| ETV1_ERF_FEV_ELF1 | 0.011 | 0.067 |
|
|
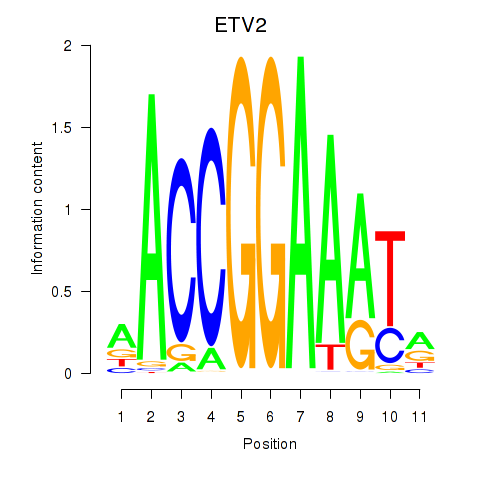
| ETV2 | 0.065 | 0.298 |
|
|
| ETV3 | -0.236 | -0.749 |
|
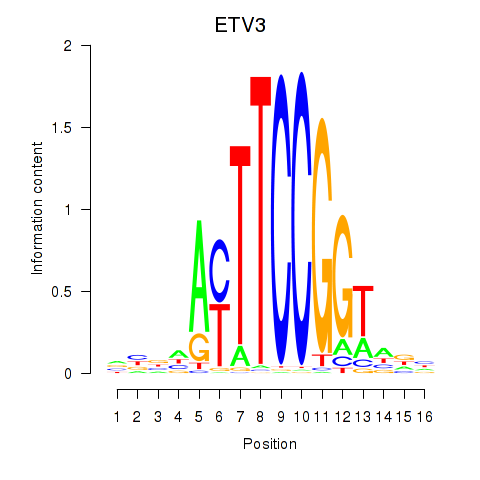
|
| ETV4_ETS2 | -0.098 | -0.431 |
|
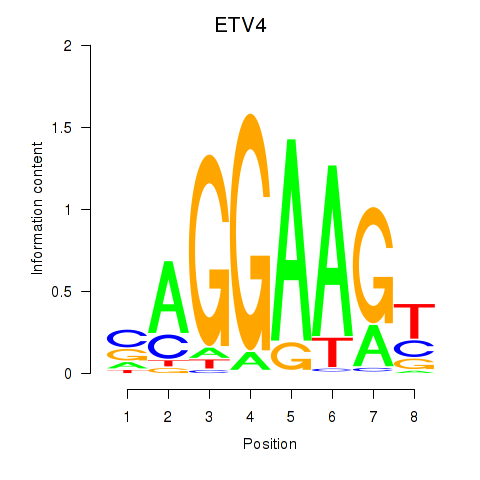
|
| ETV6 | 0.317 | 1.301 |
|
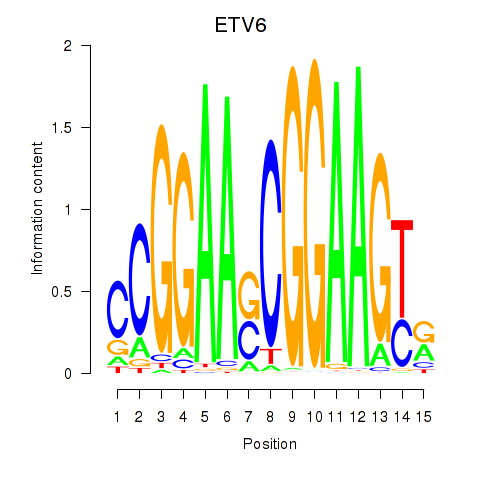
|
| ETV7 | 0.107 | 0.455 |
|
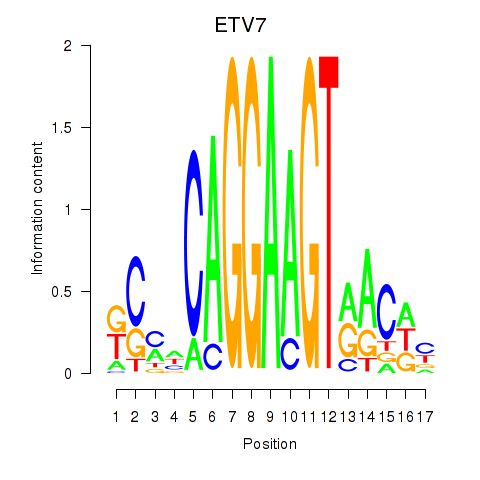
|
| EVX2 | -0.614 | -0.412 |
|
|
| EZH2 | -0.161 | -0.911 |
|
|
| FIGLA | -0.213 | -0.540 |
|
|
| FLI1 | -0.128 | -0.770 |
|
|
| FOSB | -0.265 | -0.605 |
|
|
| FOSL1 | 0.227 | 0.592 |
|
|
| FOSL2_SMARCC1 | -0.114 | -0.330 |
|
|
| FOXA1 | -0.182 | -0.395 |
|
|
| FOXA2_FOXJ3 | 0.006 | 0.019 |
|
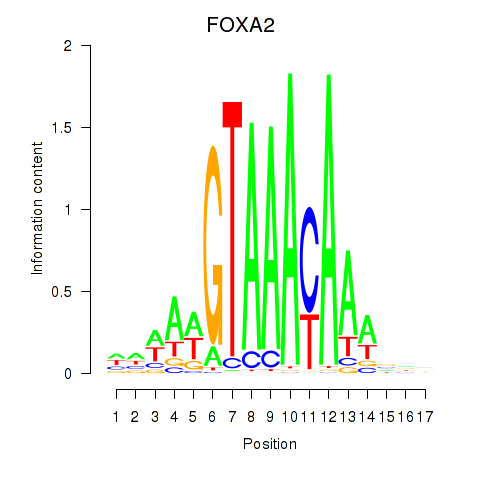
|
| FOXA3_FOXC2 | -0.091 | -0.191 |
|
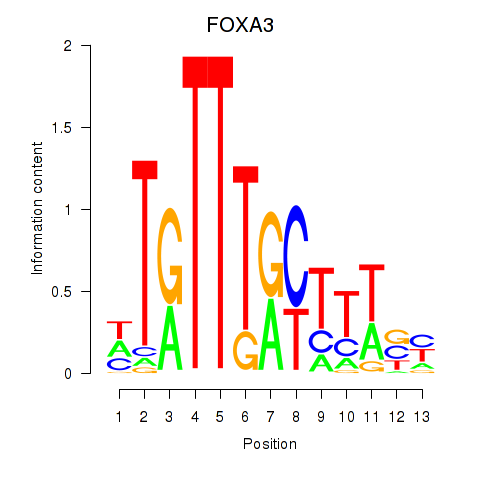
|
| FOXC1 | 0.299 | 0.736 |
|
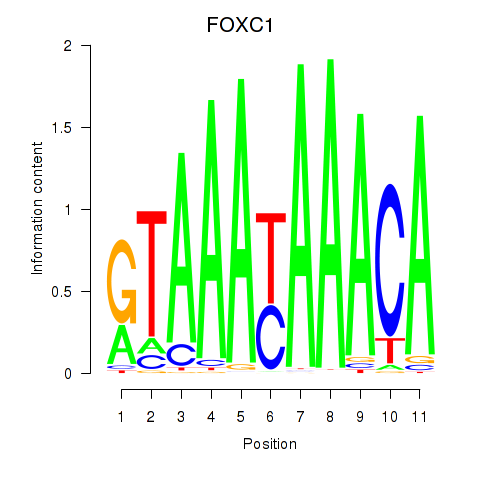
|
| FOXD1_FOXO1_FOXO6_FOXG1_FOXP1 | 0.126 | 0.450 |
|
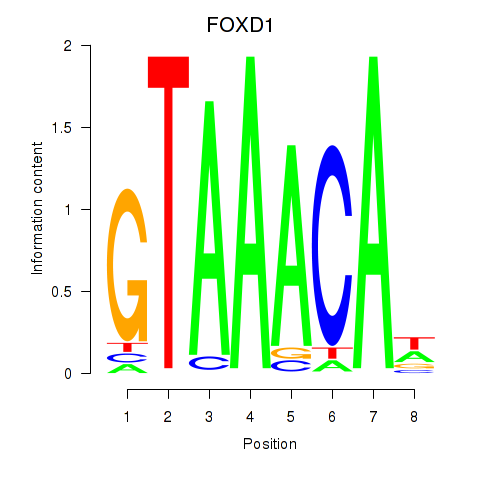
|
| FOXD3_FOXI1_FOXF1 | 0.018 | 0.071 |
|
|
| FOXF2_FOXJ1 | 0.137 | 0.298 |
|
|
| FOXJ2 | 0.113 | 0.225 |
|
|
| FOXK1_FOXP2_FOXB1_FOXP3 | 0.068 | 0.254 |
|
|
| FOXL1 | -0.150 | -0.837 |
|
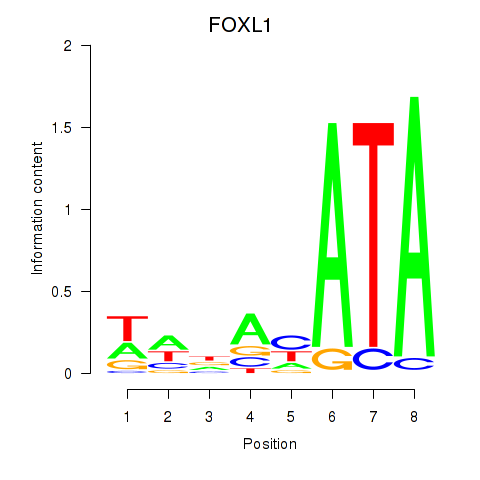
|
| FOXM1_TBL1XR1 | -0.023 | -0.339 |
|
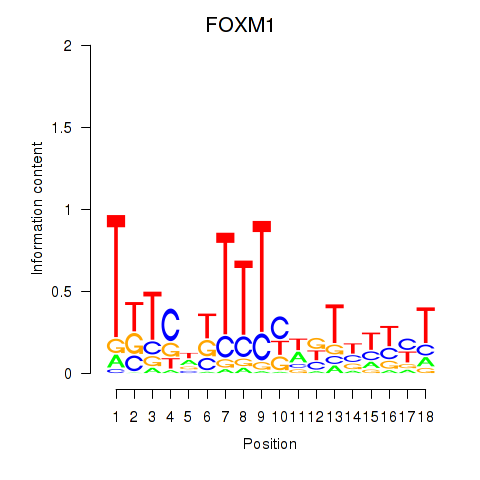
|
| FOXN1 | -0.006 | -0.016 |
|
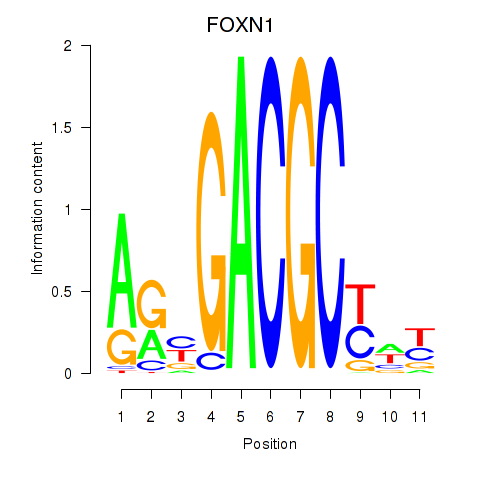
|
| FOXO3_FOXD2 | -0.363 | -0.927 |
|
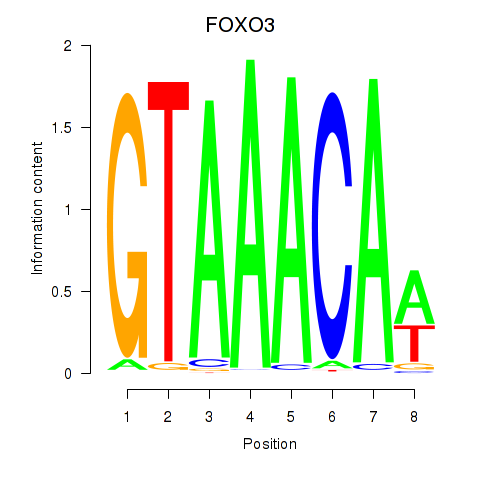
|
| FOXO4 | -0.176 | -0.465 |
|
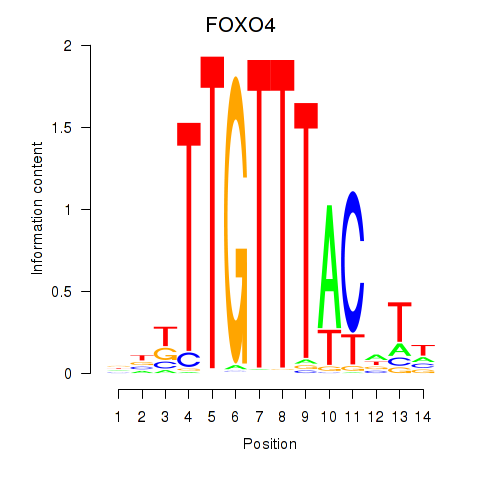
|
| FOXQ1 | 0.226 | 0.776 |
|
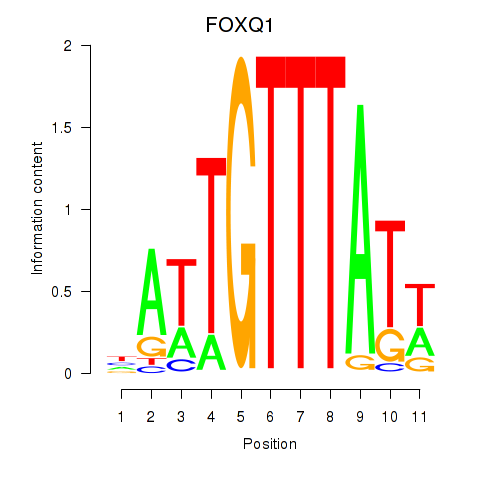
|
| FUBP1 | 0.094 | 0.179 |
|
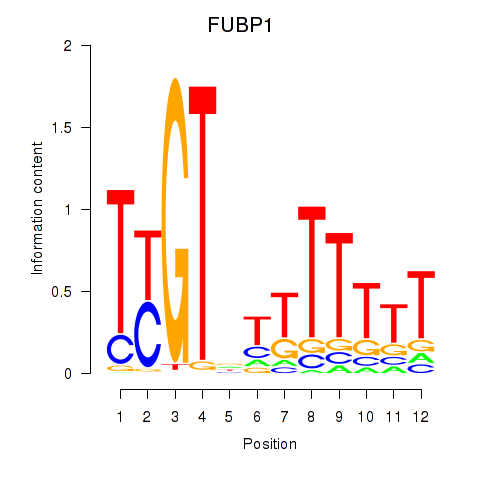
|
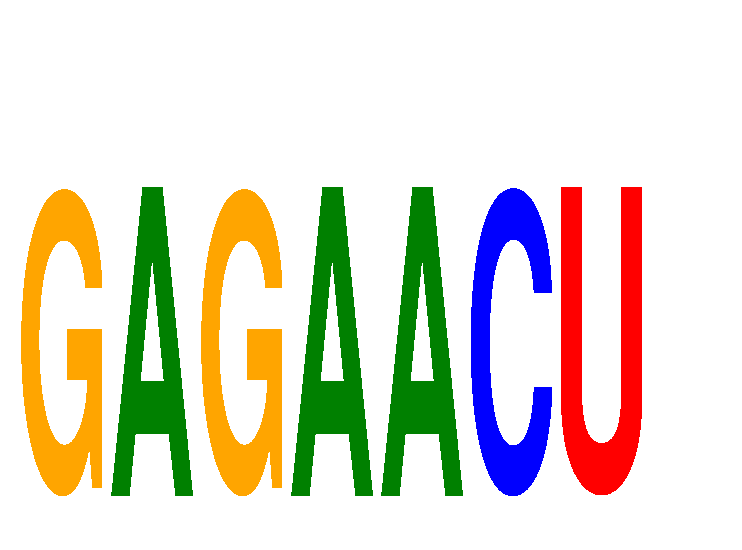
| GAGAACU | -1.705 | -0.376 |
|

|
| GAGAUGA | -0.165 | -0.166 |
|

|
| GAGGUAG | -0.282 | -0.945 |
|

|
| GATA1_GATA4 | -0.241 | -0.911 |
|
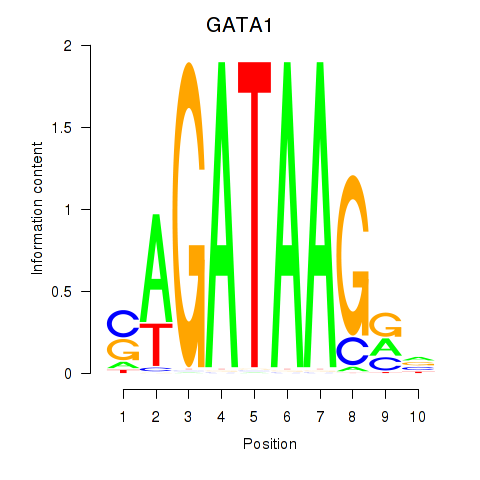
|
| GATA2 | -0.022 | -0.100 |
|
|
| GATA3 | -0.027 | -0.339 |
|
|
| GATA5 | -0.358 | -0.729 |
|
|
| GATA6 | -0.069 | -0.099 |
|
|
| GAUAUGU | -0.953 | -0.520 |
|

|
| GAUCAGA | 3.222 | 0.567 |
|

|
| GAUUGUC | -0.178 | -0.332 |
|

|
| GCAGCAU | 0.017 | 0.033 |
|

|
| GCCUGUC | -0.669 | -0.314 |
|

|
| GCM1 | -0.377 | -0.932 |
|
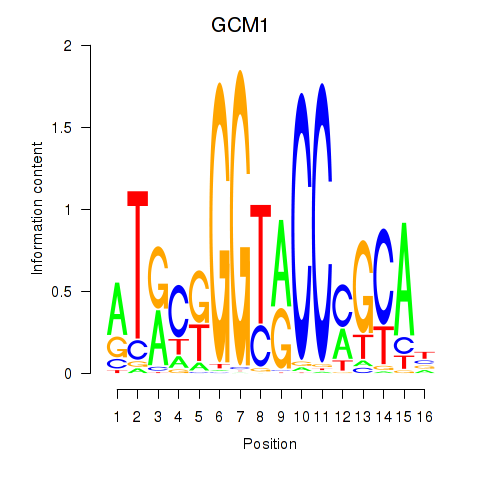
|
| GCM2 | -0.291 | -0.689 |
|
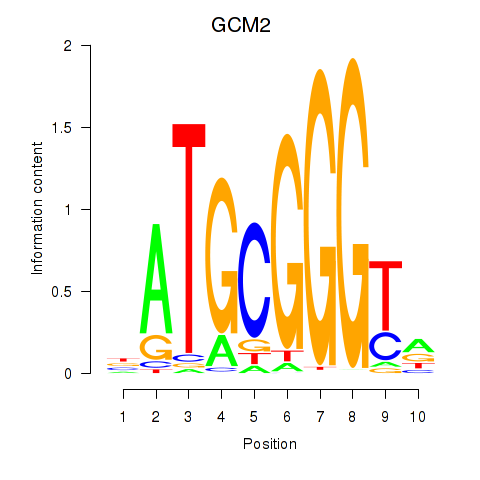
|
| GCUACAU | -0.586 | -0.795 |
|

|
| GCUGGUG | -0.177 | -0.384 |
|

|
| GFI1 | -0.237 | -0.904 |
|
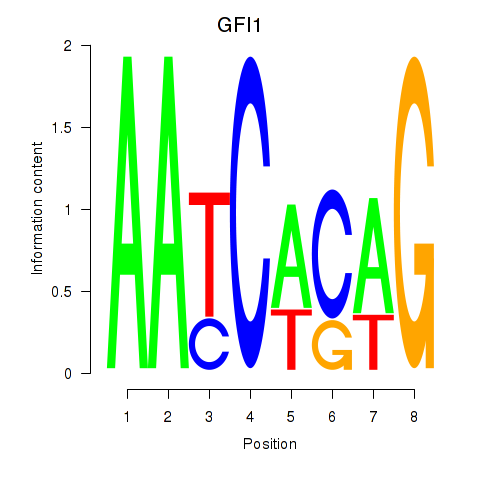
|
| GFI1B | -0.162 | -0.501 |
|
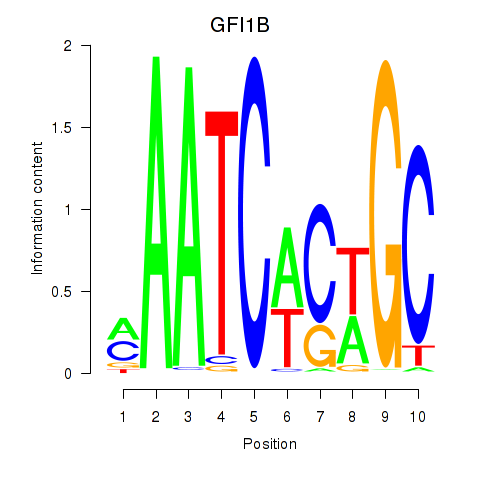
|
| GGAAGAC | 0.494 | 0.742 |
|

|
| GGAAUGU | 0.660 | 1.671 |
|

|
| GGACGGA | -1.670 | -0.472 |
|

|
| GGAGUGU | 0.117 | 0.061 |
|

|
| GGCAAGA | 0.099 | 0.104 |
|

|
| GGCAGUG | -0.170 | -0.392 |
|

|
| GGCUCAG | -0.409 | -0.707 |
|

|
| GLI2 | 0.002 | 0.009 |
|
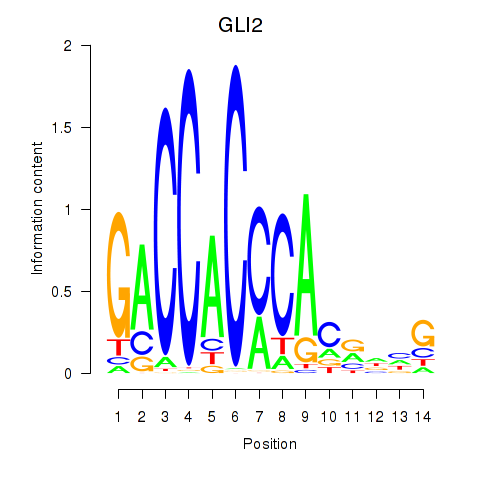
|
| GLI3 | 0.010 | 0.058 |
|
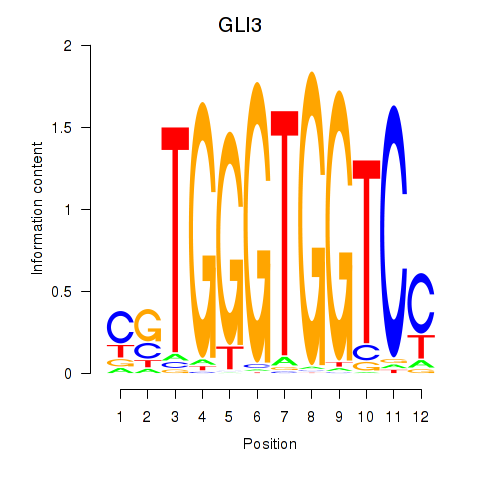
|
| GLIS1 | -0.906 | -0.518 |
|
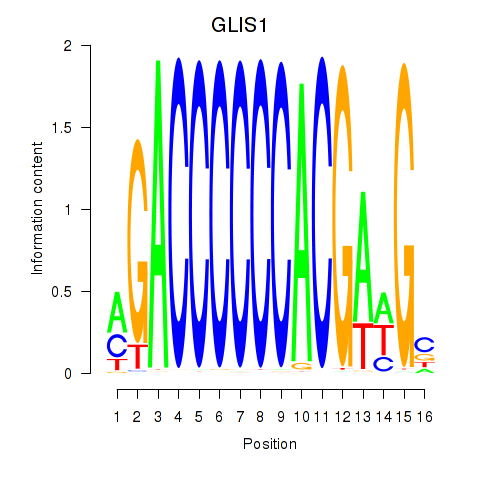
|
| GLIS2 | 0.099 | 0.311 |
|
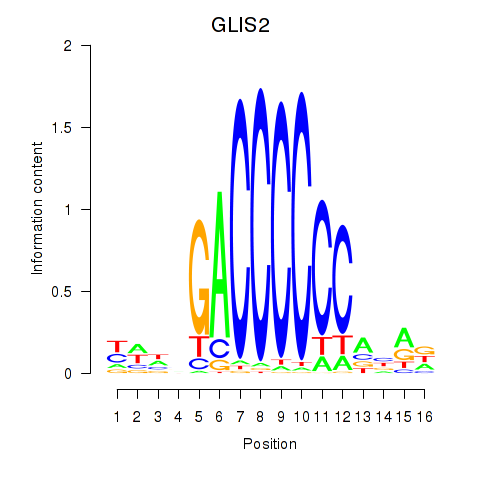
|
| GLIS3 | -0.072 | -0.094 |
|
|
| GMEB1 | 0.028 | 0.178 |
|
|
| GMEB2 | 0.518 | 2.497 |
|
|
| GRHL1 | -0.299 | -0.377 |
|
|
| GSC_GSC2 | -0.467 | -0.836 |
|
|
| GTF2I | 0.036 | 0.299 |
|
|
| GUAAACA | -0.021 | -0.072 |
|

|
| GUAACAG | 0.529 | 0.620 |
|

|
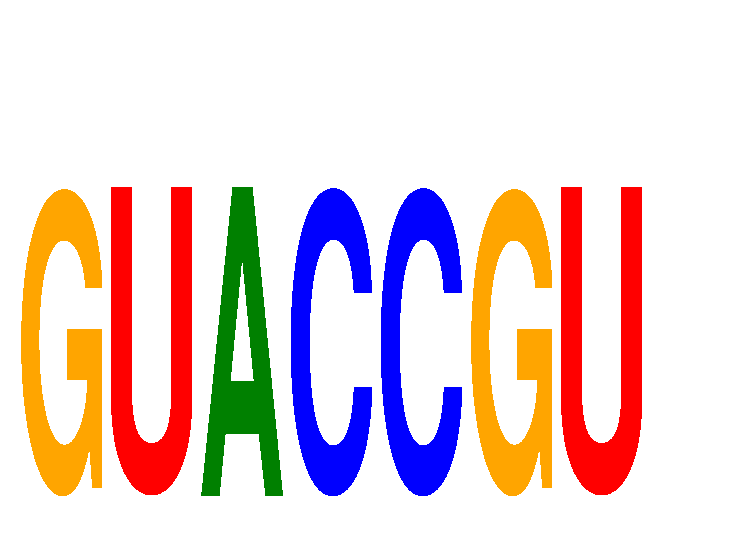
| GUACCGU | 0.218 | 0.019 |
|

|
| GUAGUGU | 0.070 | 0.111 |
|

|
| GUCAGUU | 0.110 | 0.111 |
|

|
| GUGCAAA | 0.176 | 0.486 |
|

|
| GZF1 | 0.351 | 0.408 |
|
|
| HAND1 | -0.528 | -0.865 |
|
|
| HBP1 | -0.239 | -0.653 |
|
|
| HDX | -1.082 | -0.856 |
|
|
| HES1 | 0.114 | 0.555 |
|
|
| HES7_HES5 | 0.015 | 0.036 |
|
|
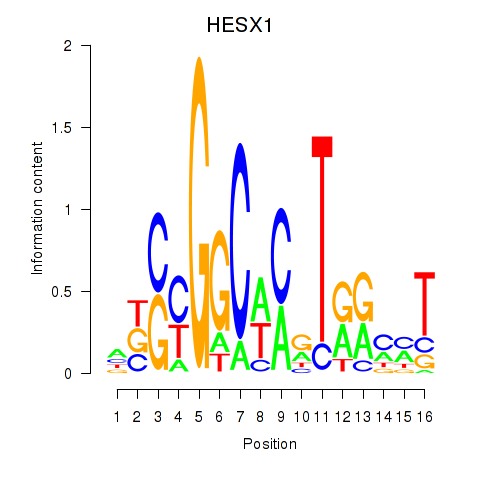
| HESX1 | -0.315 | -0.619 |
|
|
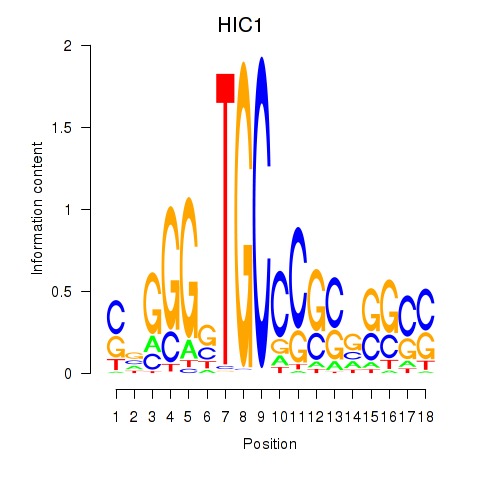
| HIC1 | -0.092 | -0.633 |
|
|
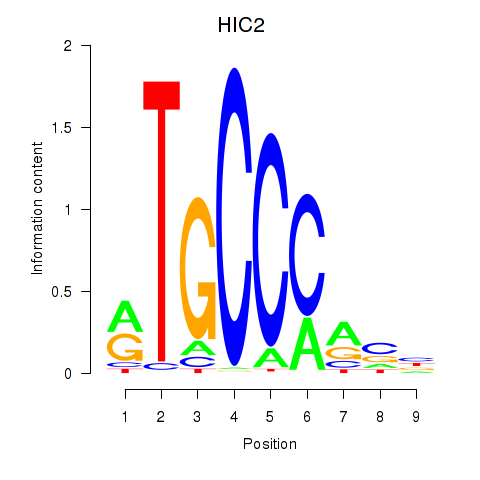
| HIC2 | -0.198 | -1.488 |
|
|
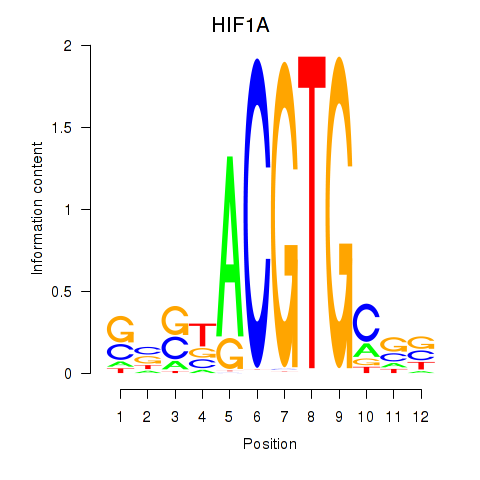
| HIF1A | 0.188 | 0.502 |
|
|
| HINFP | -0.005 | -0.014 |
|
|
| HINFP1 | 0.606 | 0.598 |
|
|
| HIVEP1 | -0.234 | -0.800 |
|
|
| HLF_TEF | 0.372 | 0.787 |
|
|
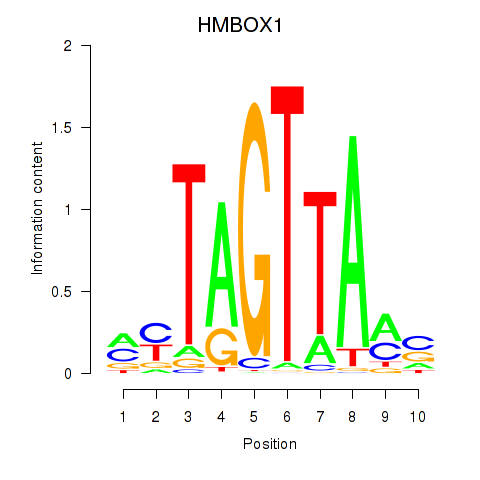
| HMBOX1 | 0.256 | 0.596 |
|
|
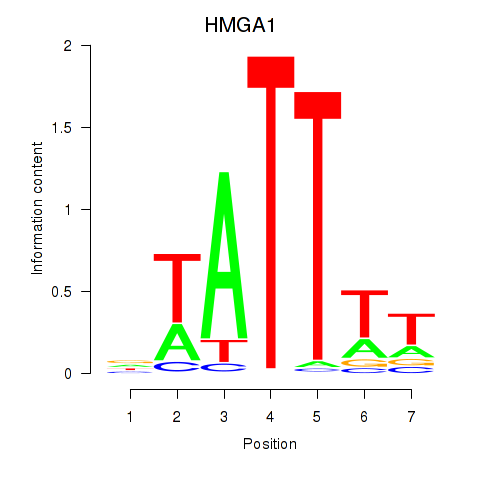
| HMGA1 | 0.092 | 0.368 |
|
|
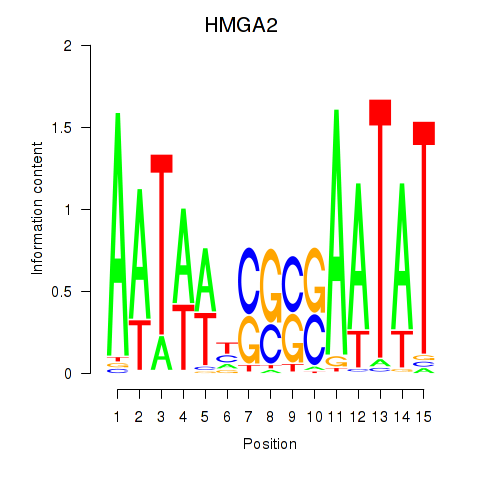
| HMGA2 | 0.409 | 0.555 |
|
|
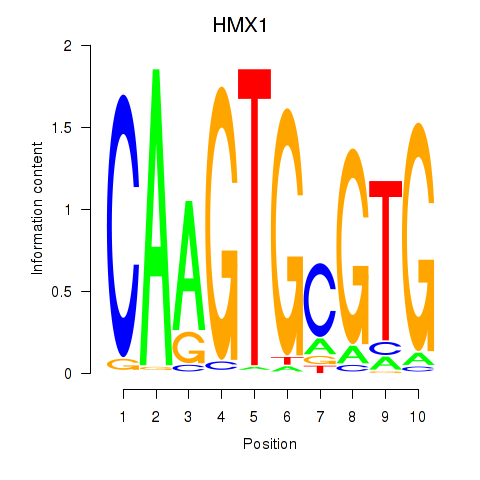
| HMX1 | -0.059 | -0.175 |
|
|
| HMX2 | 0.234 | 0.446 |
|
|
| HMX3 | -0.331 | -1.355 |
|
|
| HNF1A_HNF1B | -0.475 | -1.135 |
|
|
| HNF4A | -0.202 | -0.437 |
|
|
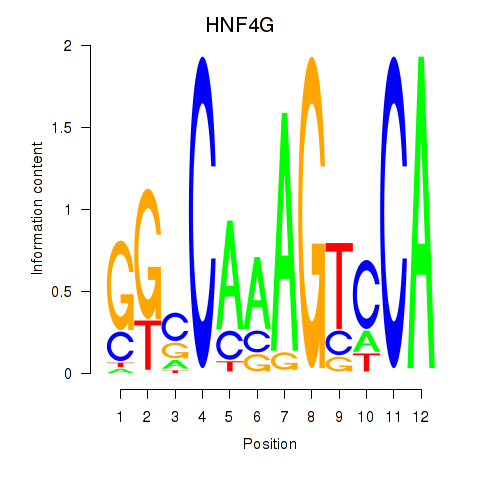
| HNF4G | -0.010 | -0.041 |
|
|
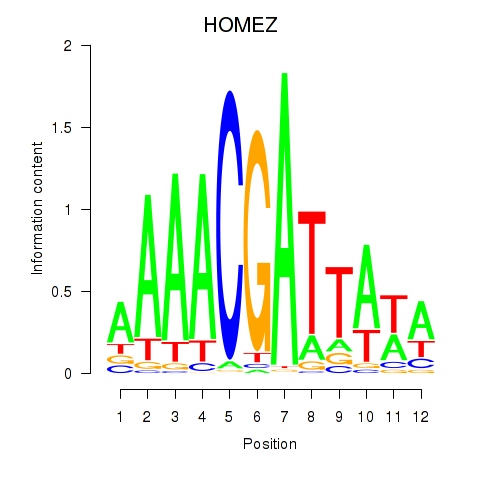
| HOMEZ | 0.176 | 0.685 |
|
|
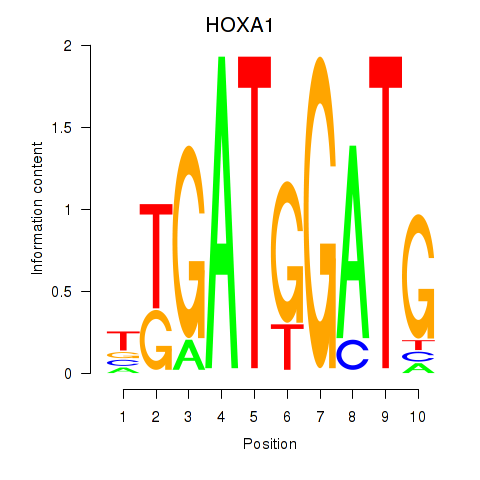
| HOXA1 | -0.079 | -0.225 |
|
|
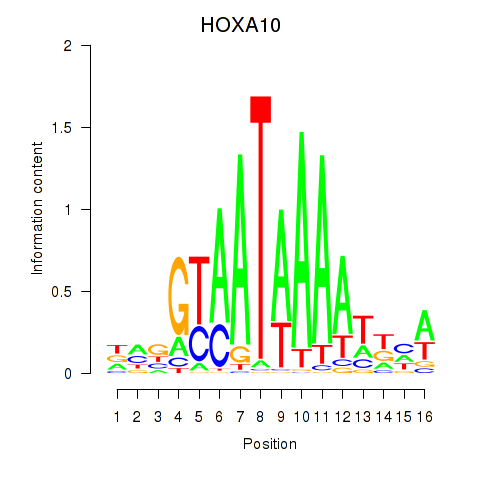
| HOXA10_HOXB9 | -0.516 | -0.949 |
|
|
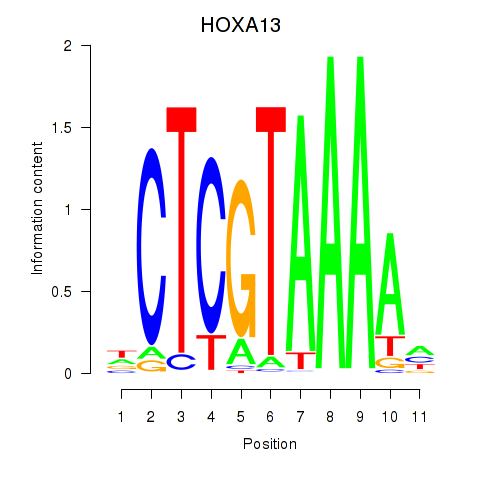
| HOXA13 | -0.202 | -0.492 |
|
|
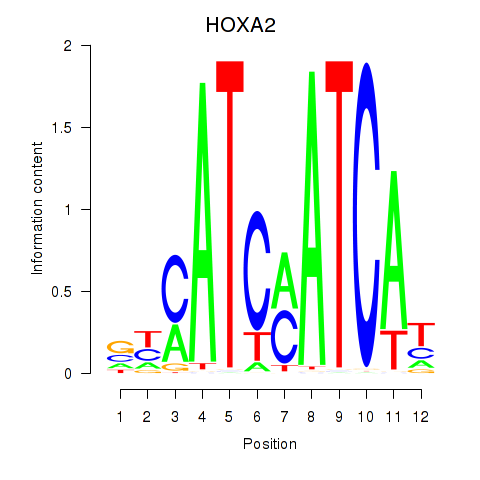
| HOXA2_HOXB1 | -0.497 | -0.854 |
|
|
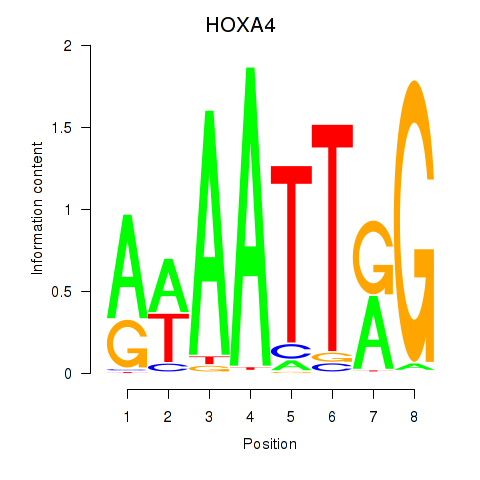
| HOXA4 | -0.113 | -0.198 |
|
|
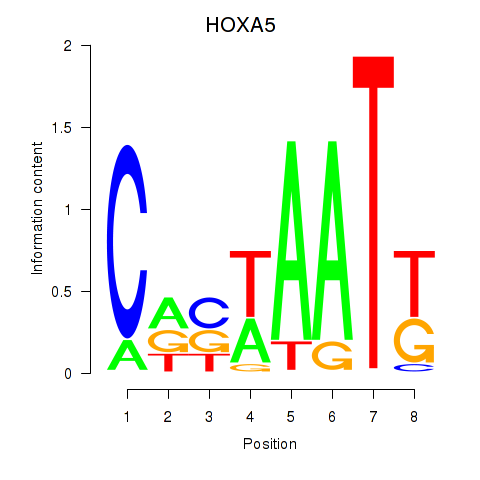
| HOXA5 | -0.266 | -0.430 |
|
|
| HOXA6 | -0.086 | -0.140 |
|
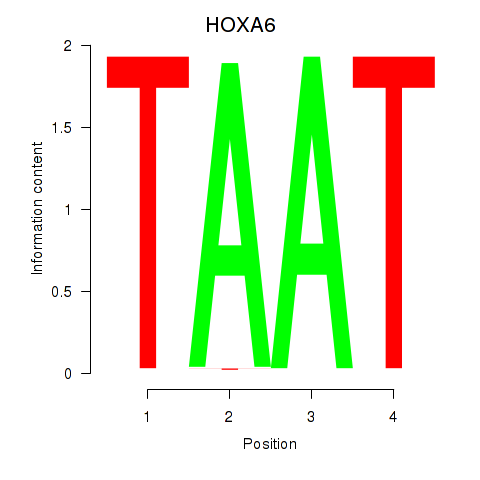
|
| HOXA9 | 0.006 | 0.017 |
|
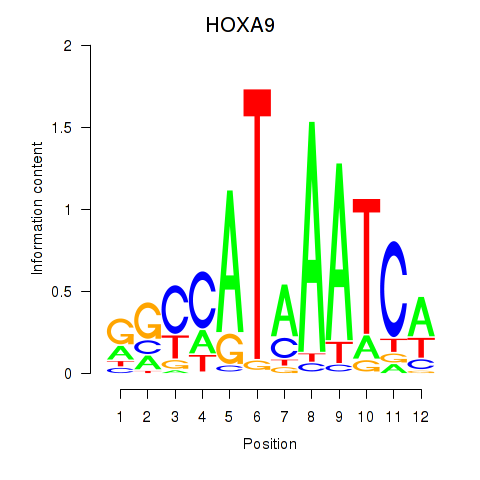
|
| HOXB13 | -0.038 | -0.042 |
|
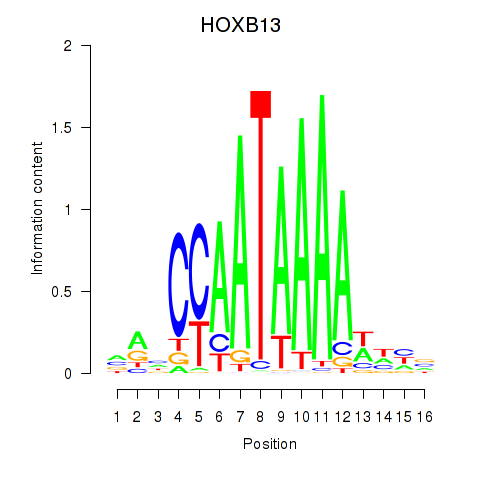
|
| HOXB2_UNCX_HOXD3 | 0.182 | 0.595 |
|
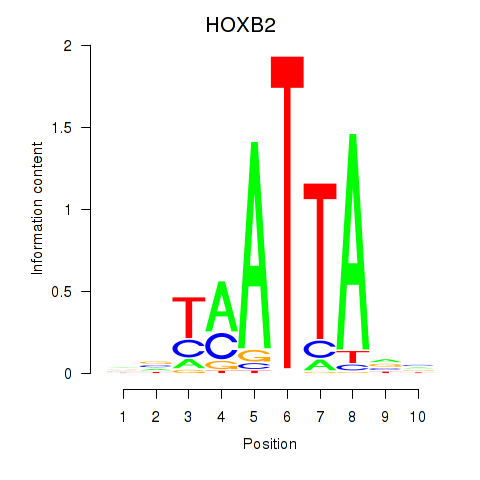
|
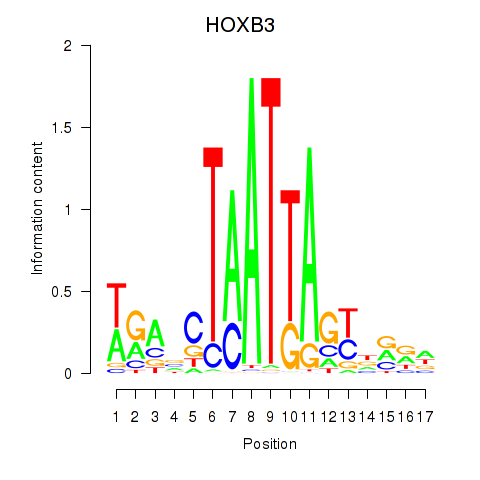
| HOXB3 | -0.052 | -0.114 |
|
|
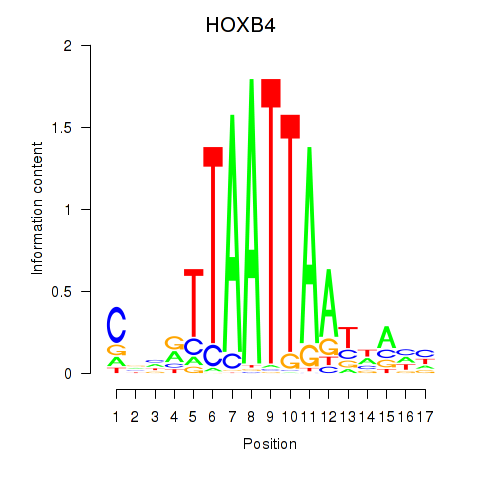
| HOXB4_LHX9 | 0.118 | 0.241 |
|
|
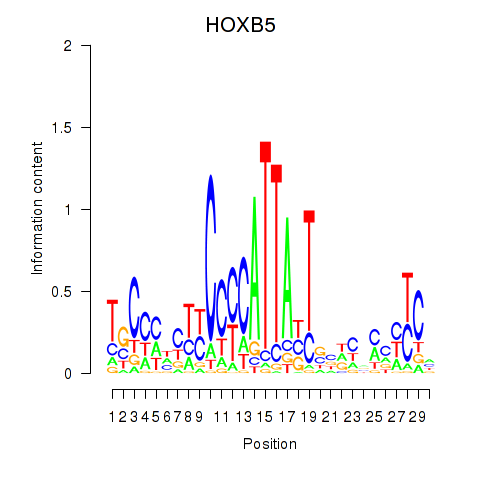
| HOXB5 | -0.196 | -0.274 |
|
|
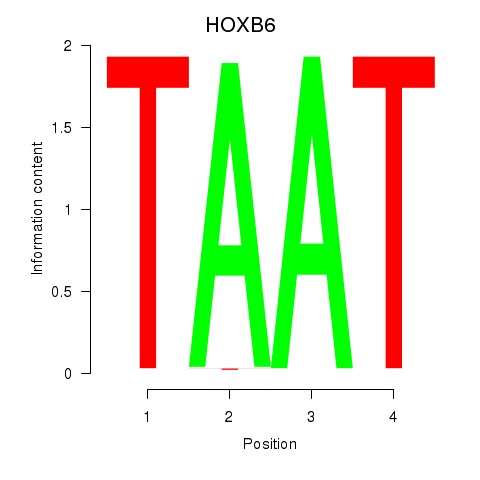
| HOXB6_PRRX2 | -0.183 | -0.733 |
|
|
| HOXB7 | -0.375 | -0.955 |
|
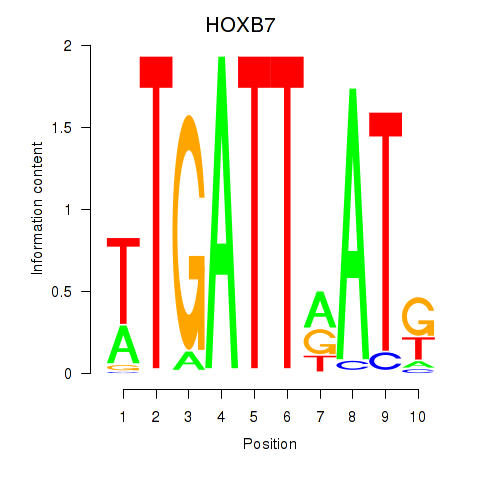
|
| HOXB8 | 0.123 | 0.500 |
|
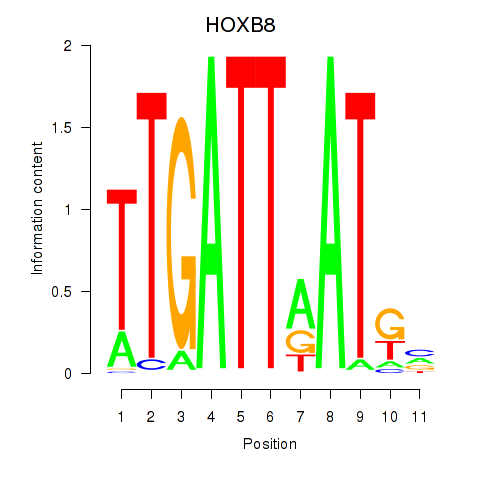
|
| HOXC10_HOXD13 | 0.038 | 0.121 |
|
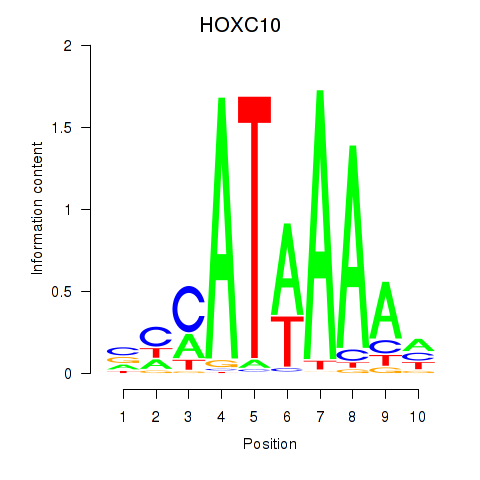
|
| HOXC11 | 0.022 | 0.038 |
|
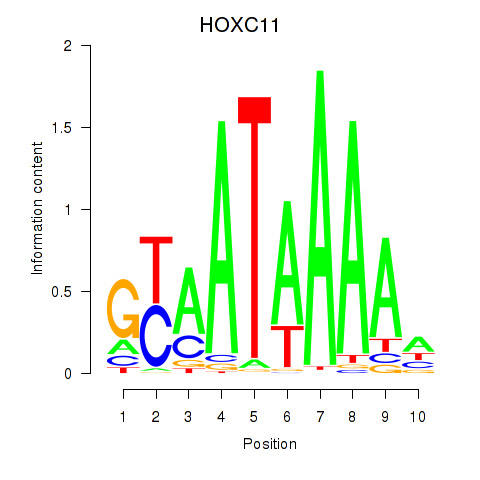
|
| HOXC12_HOXD12 | 0.199 | 0.559 |
|
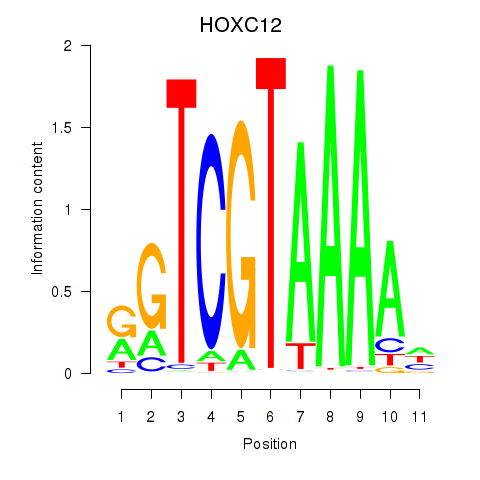
|
| HOXC13 | -0.317 | -0.686 |
|
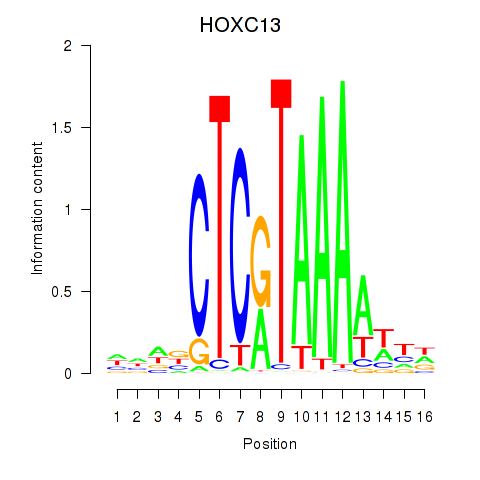
|
| HOXC6_HOXA7 | -0.235 | -0.324 |
|
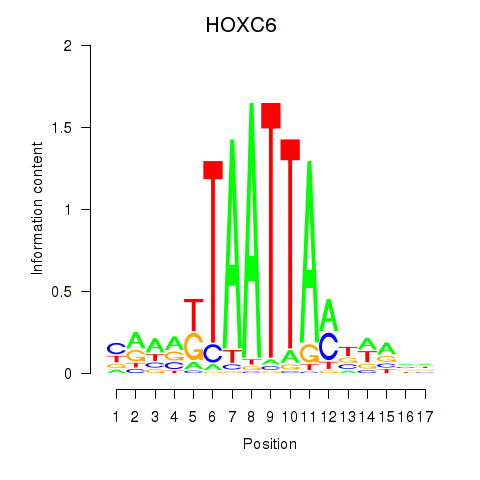
|
| HOXC8 | 0.157 | 0.933 |
|
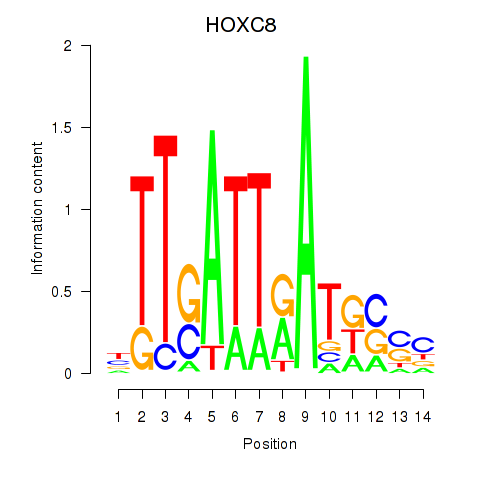
|
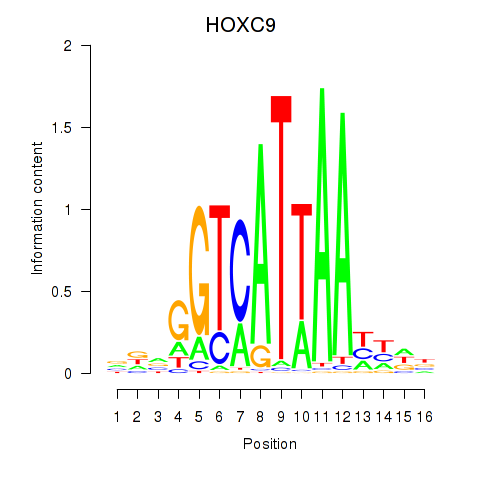
| HOXC9 | 0.152 | 0.370 |
|
|
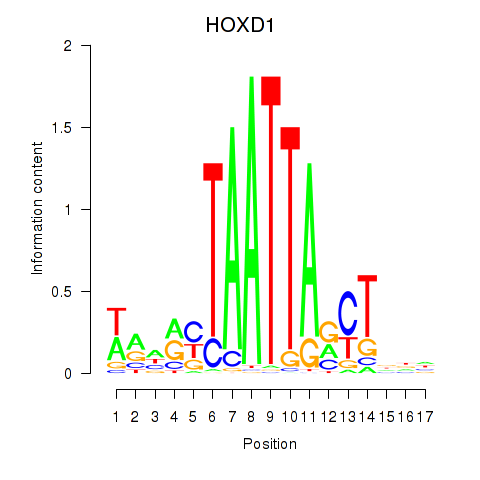
| HOXD1 | 0.989 | 0.932 |
|
|
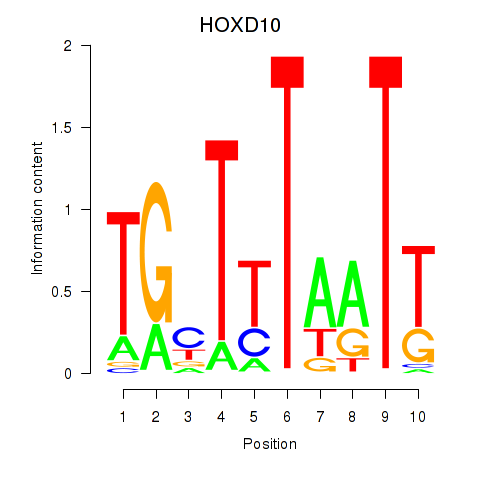
| HOXD10 | -0.044 | -0.126 |
|
|
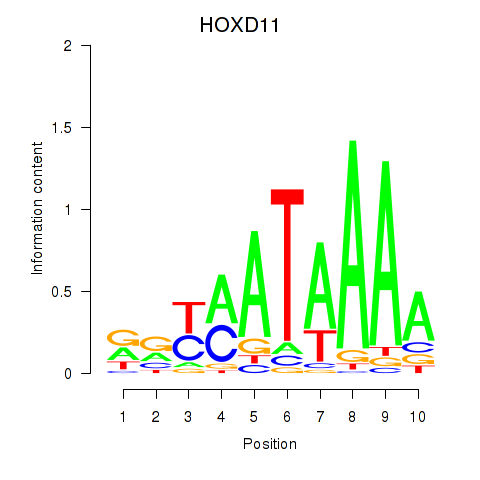
| HOXD11_HOXA11 | 0.081 | 0.239 |
|
|
| HOXD4 | -0.098 | -0.177 |
|
|
| HOXD9 | -0.182 | -0.409 |
|
|
| HSF1 | -0.067 | -0.272 |
|
|
| HSF2 | 0.119 | 0.227 |
|
|
| HSF4 | 0.692 | 1.762 |
|
|
| HSFY2 | 0.064 | 0.252 |
|
|
| ID4_TCF4_SNAI2 | -0.169 | -1.584 |
|
|
| IKZF1 | -0.007 | -0.136 |
|
|
| IKZF2 | 0.286 | 0.892 |
|
|
| INSM1 | 0.003 | 0.008 |
|

|
| IRF2_STAT2_IRF8_IRF1 | 0.160 | 0.661 |
|
|
| IRF3 | -0.060 | -0.133 |
|
|
| IRF6_IRF4_IRF5 | 0.204 | 0.460 |
|
|
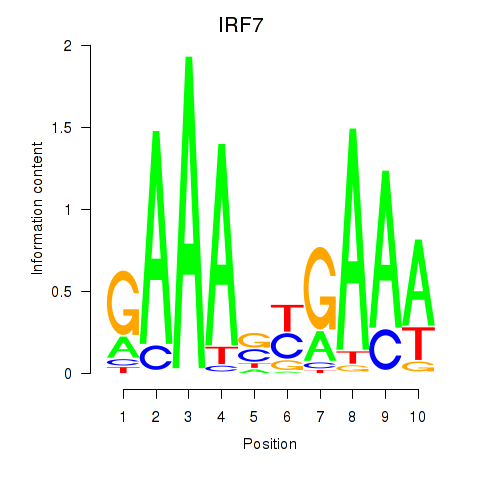
| IRF7 | -0.027 | -0.114 |
|
|
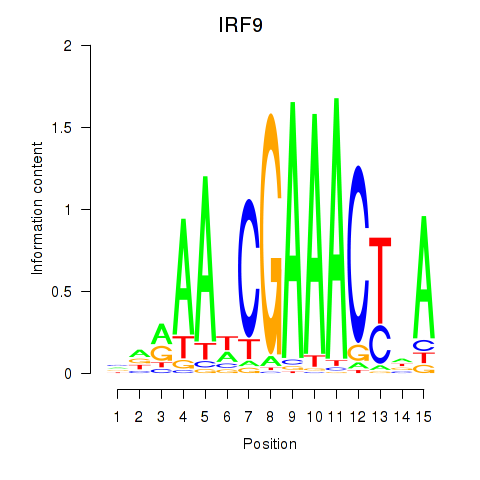
| IRF9 | -0.025 | -0.038 |
|
|
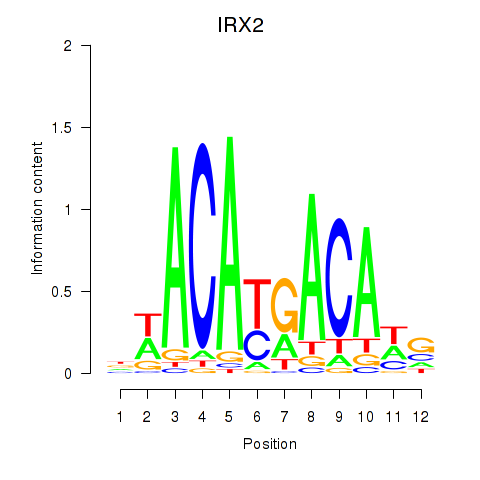
| IRX2 | 0.091 | 0.165 |
|
|
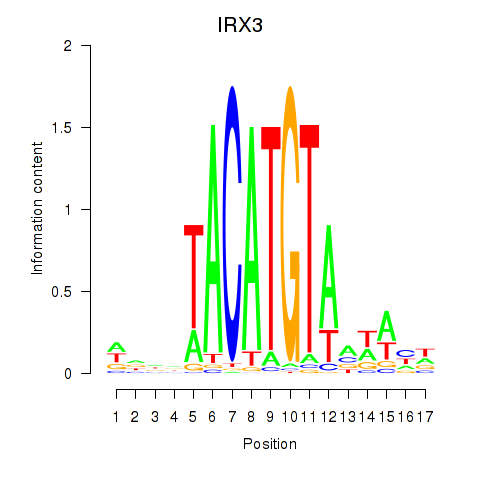
| IRX3 | -0.308 | -1.142 |
|
|
| IRX5 | 0.439 | 0.769 |
|
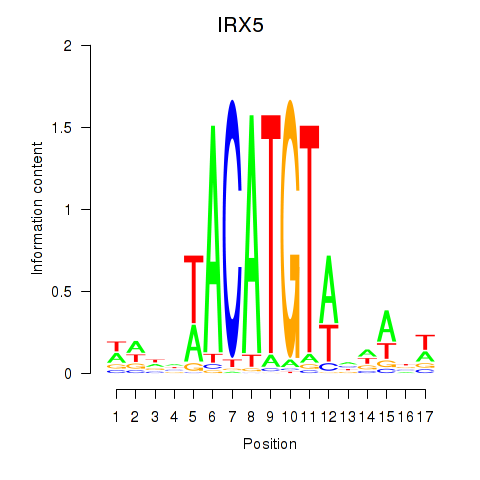
|
| IRX6_IRX4 | 0.107 | 0.096 |
|
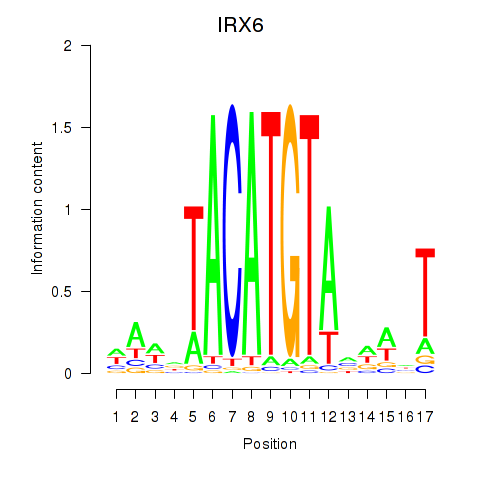
|
| ISL1 | -0.208 | -0.458 |
|
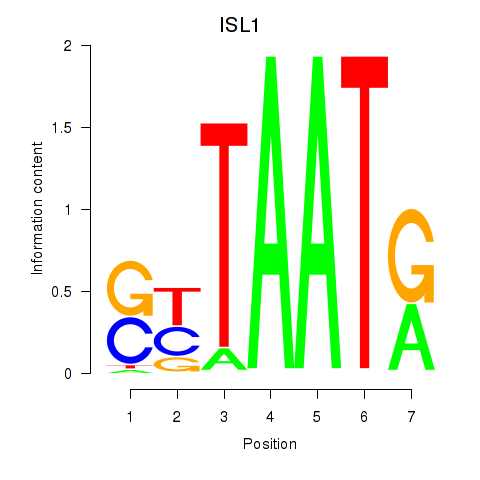
|
| ISL2 | -0.143 | -0.218 |
|
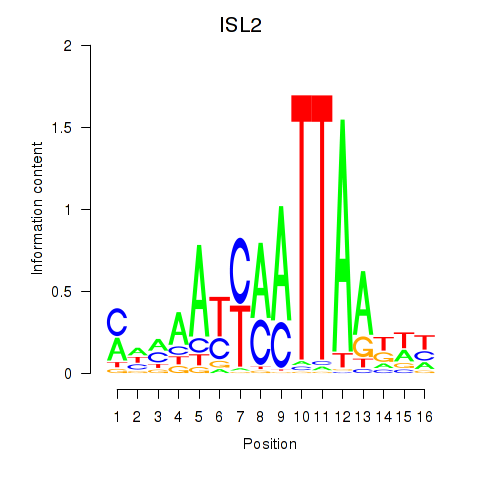
|
| ISX | 0.028 | 0.052 |
|
|
| JDP2 | -0.266 | -0.556 |
|
|
| JUN | -0.066 | -0.219 |
|
|
| JUND | -0.256 | -0.479 |
|
|
| KLF1 | 0.489 | 1.785 |
|
|
| KLF12 | 0.137 | 0.476 |
|
|
| KLF13 | -0.310 | -0.816 |
|
|
| KLF14_SP8 | -0.061 | -0.521 |
|
|
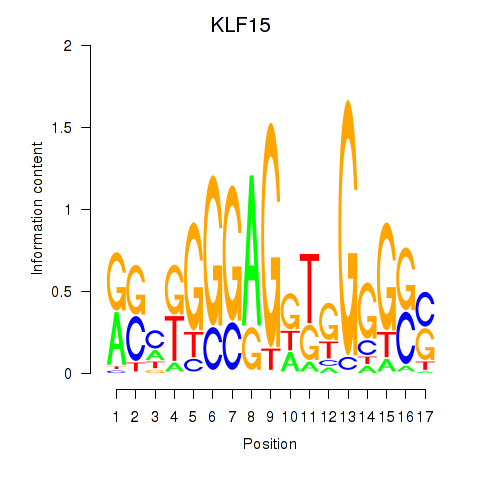
| KLF15 | 0.028 | 0.090 |
|
|
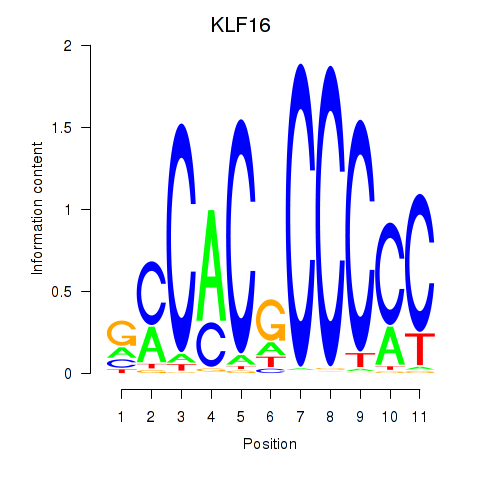
| KLF16_SP2 | 0.030 | 0.398 |
|
|
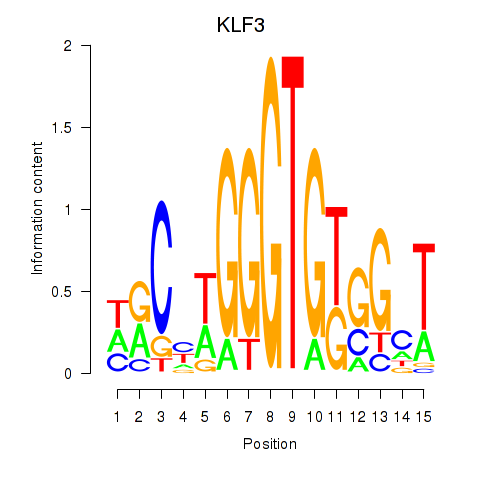
| KLF3 | -0.193 | -0.763 |
|
|
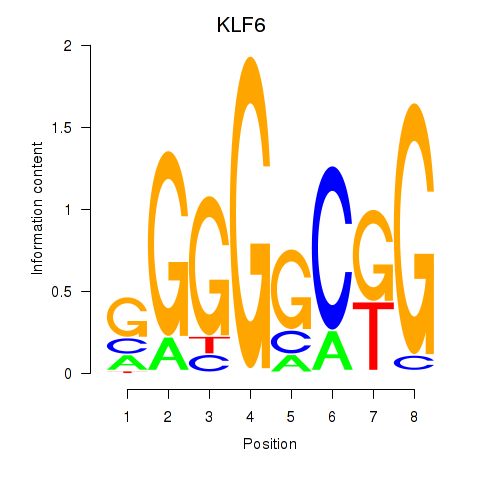
| KLF6 | -0.160 | -0.585 |
|
|
| KLF7 | -0.383 | -0.401 |
|
|
| KLF8 | -0.074 | -0.794 |
|
|
| LEF1 | -0.191 | -0.697 |
|
|
| LHX2 | 0.193 | 0.325 |
|
|
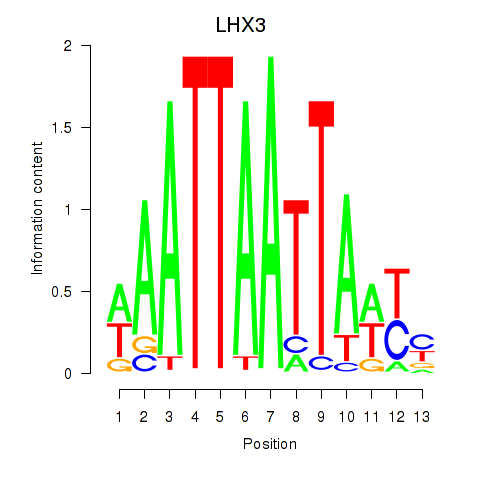
| LHX3 | 0.478 | 0.733 |
|
|
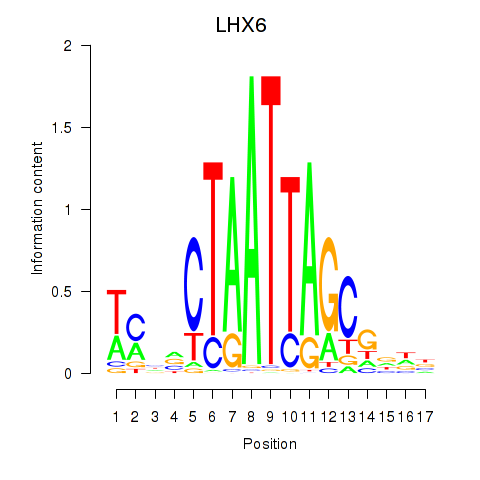
| LHX6 | -0.251 | -0.556 |
|
|
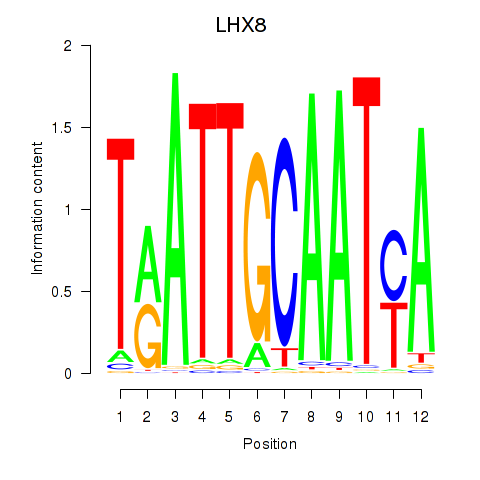
| LHX8 | 0.002 | 0.005 |
|
|
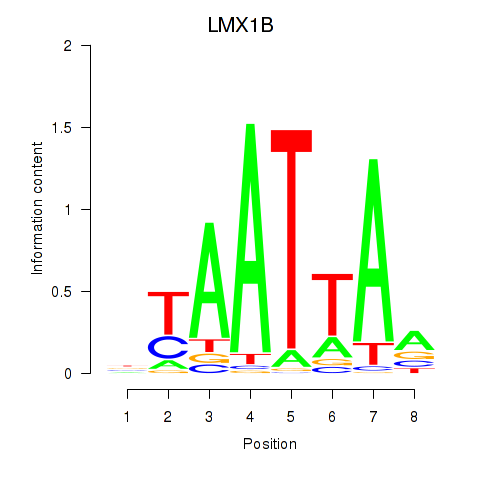
| LMX1B_MNX1_RAX2 | 0.064 | 0.101 |
|
|
| MAFA | 0.151 | 0.261 |
|
|
| MAFB | 0.078 | 0.633 |
|
|
| MAFF_MAFG | 0.036 | 0.115 |
|
|
| MAFK | 0.016 | 0.092 |
|
|
| MAF_NRL | -0.234 | -0.886 |
|
|
| MAX_TFEB | 0.335 | 1.060 |
|
|
| MAZ_ZNF281_GTF2F1 | -0.090 | -1.234 |
|
|
| MBD2 | -0.370 | -1.298 |
|
|
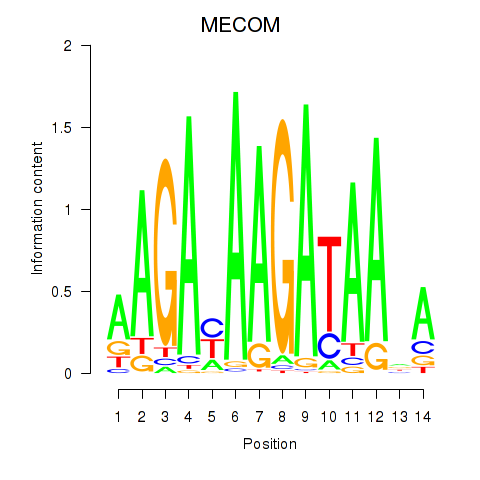
| MECOM | -0.367 | -0.847 |
|
|
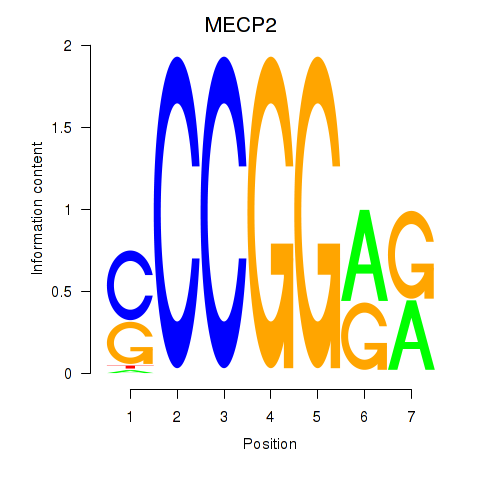
| MECP2 | 0.041 | 0.746 |
|
|
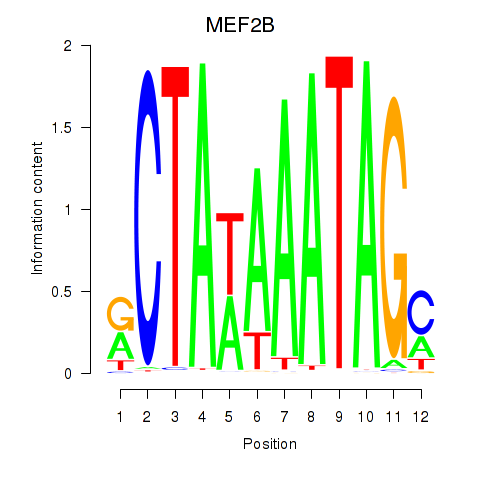
| MEF2B | -0.261 | -0.416 |
|
|
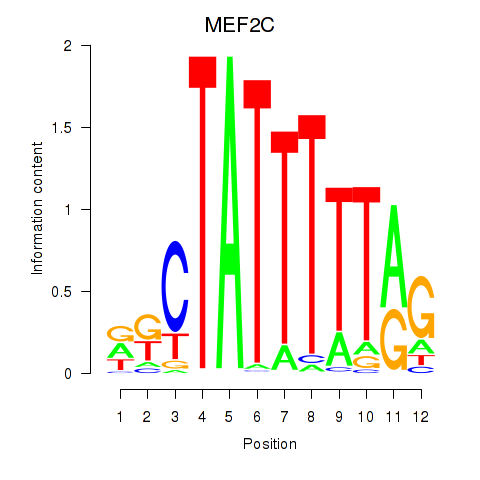
| MEF2C | 0.118 | 0.234 |
|
|
| MEF2D_MEF2A | -0.017 | -0.035 |
|
|
| MEIS1 | -0.039 | -0.185 |
|
|
| MEIS2 | -0.039 | -0.388 |
|
|
| MEIS3_TGIF2LX | -0.233 | -0.511 |
|
|
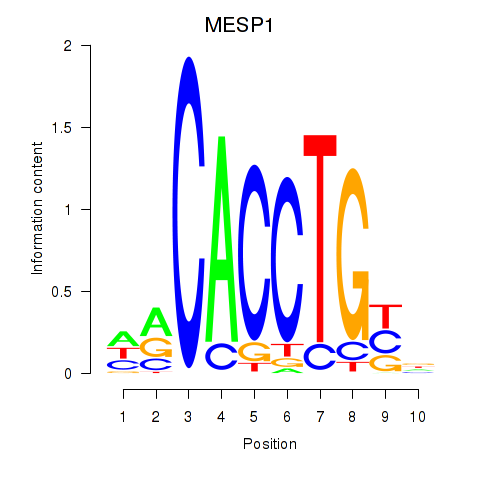
| MESP1 | 0.107 | 0.476 |
|
|
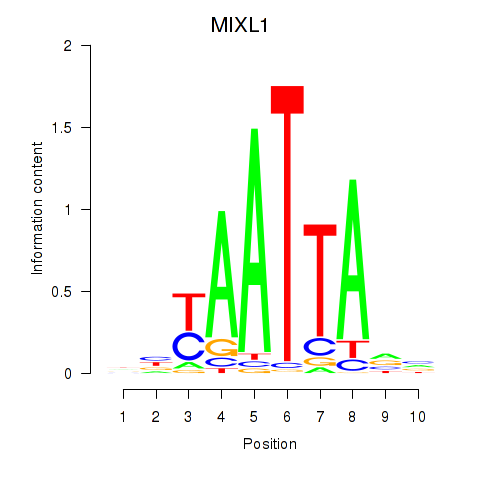
| MIXL1_GSX1_BSX_MEOX2_LHX4 | 0.057 | 0.170 |
|
|
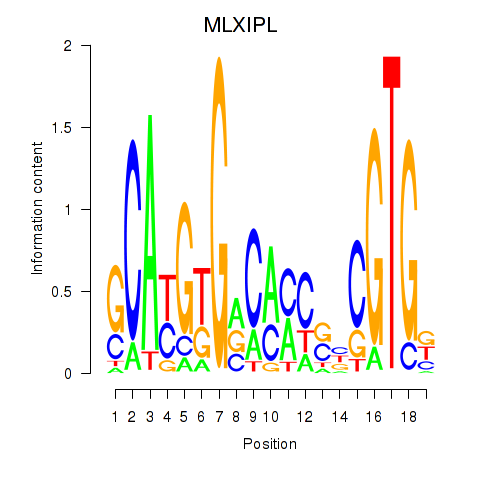
| MLXIPL | -0.087 | -0.751 |
|
|
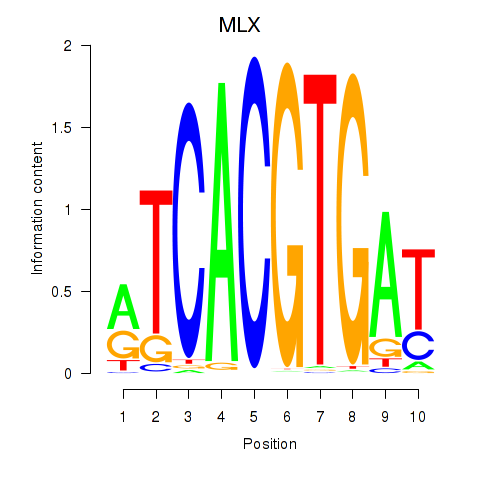
| MLX_USF2_USF1_PAX2 | -0.023 | -0.086 |
|
|
| MNT_HEY1_HEY2 | -0.251 | -1.316 |
|
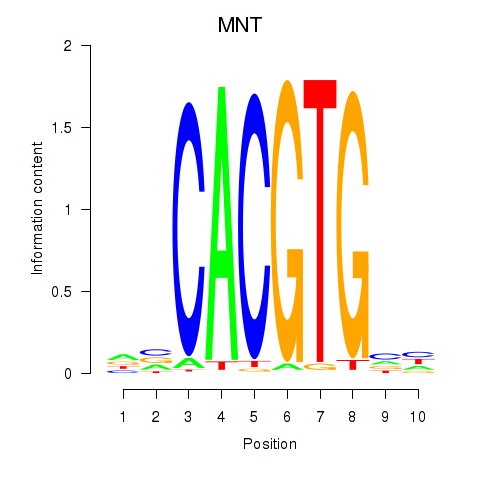
|
| MSX1 | -0.277 | -0.525 |
|
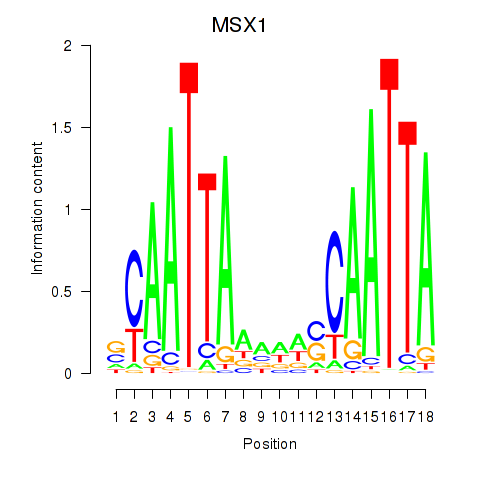
|
| MSX2 | -0.139 | -0.198 |
|
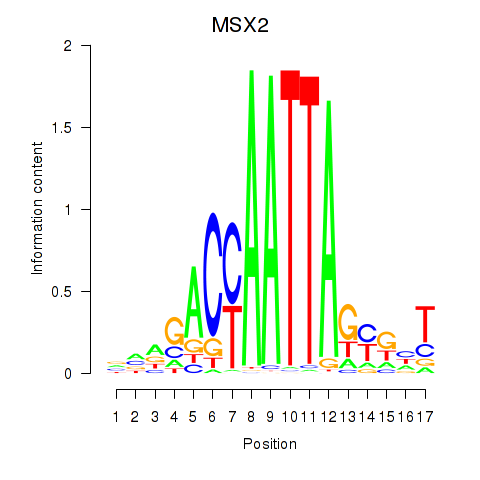
|
| MXI1_MYC_MYCN | 0.185 | 1.303 |
|
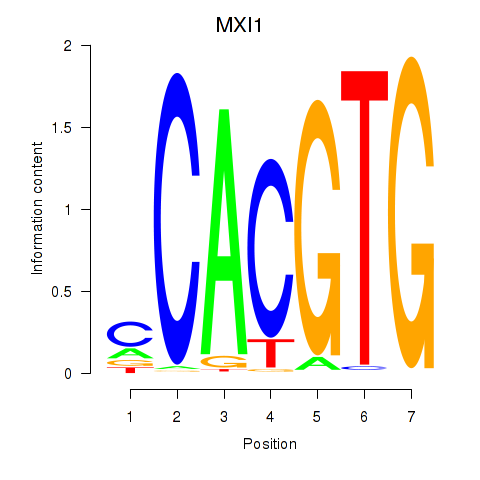
|
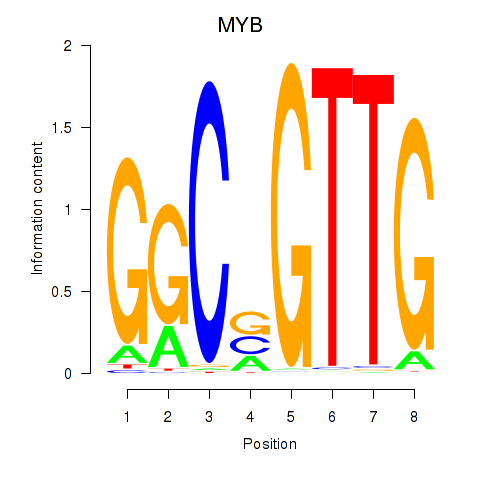
| MYB | 0.182 | 0.998 |
|
|
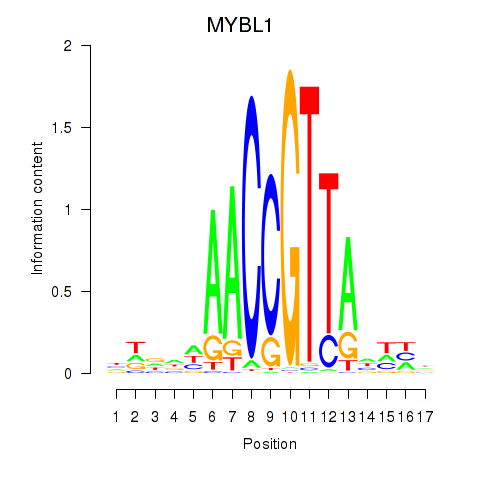
| MYBL1 | -0.025 | -0.076 |
|
|
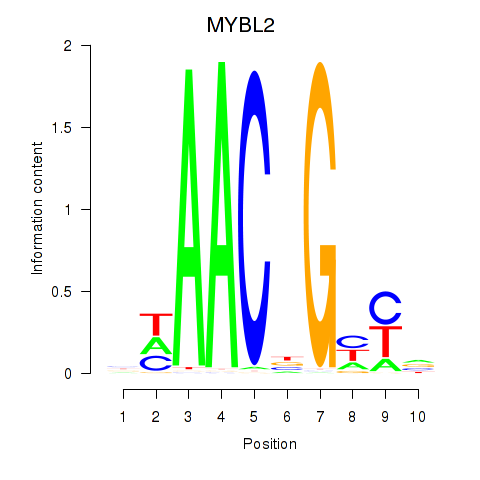
| MYBL2 | 0.035 | 0.145 |
|
|
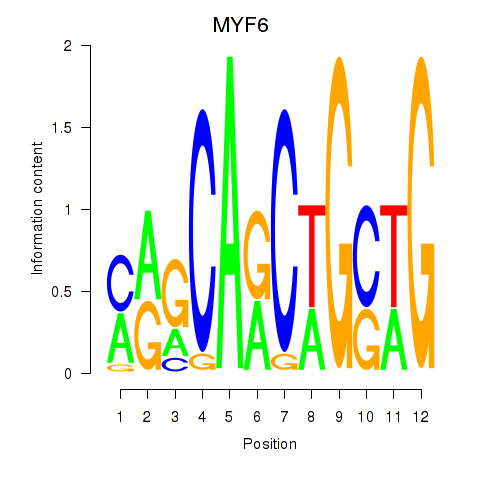
| MYF6 | -0.005 | -0.027 |
|
|
| MYOD1 | -0.104 | -0.358 |
|
|
| MZF1 | 0.013 | 0.069 |
|
|
| NANOG | 0.419 | 0.836 |
|
|
| NFAT5 | -0.252 | -0.504 |
|
|
| NFATC1 | 0.122 | 0.382 |
|
|
| NFATC2_NFATC3 | -0.049 | -0.195 |
|
|
| NFATC4 | -0.293 | -0.911 |
|
|
| NFE2L1 | -0.163 | -0.568 |
|
|
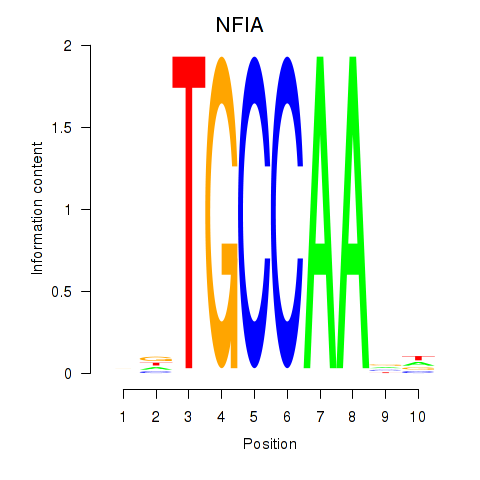
| NFIA | -0.142 | -0.509 |
|
|
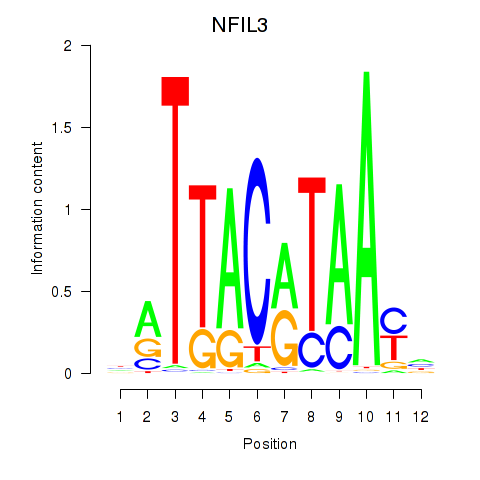
| NFIL3 | -0.044 | -0.239 |
|
|
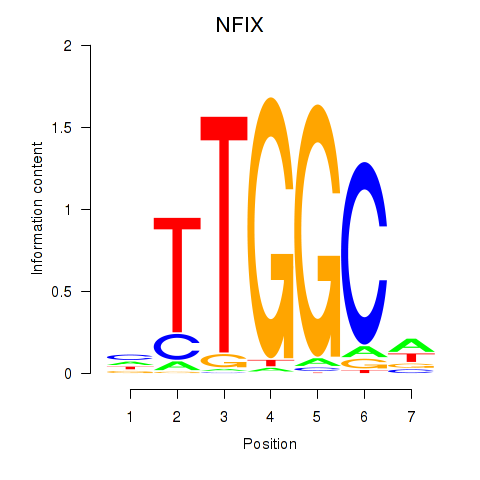
| NFIX_NFIB | -0.068 | -0.695 |
|
|
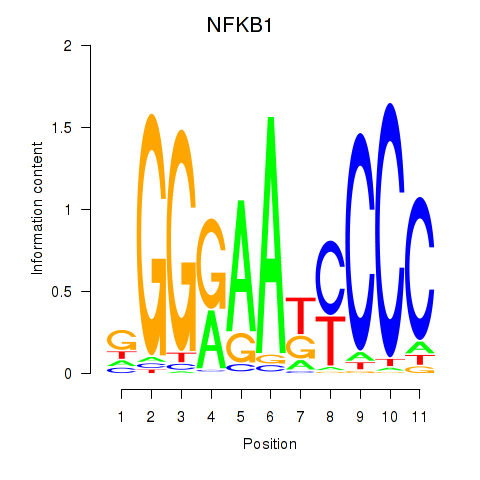
| NFKB1 | -0.255 | -0.623 |
|
|
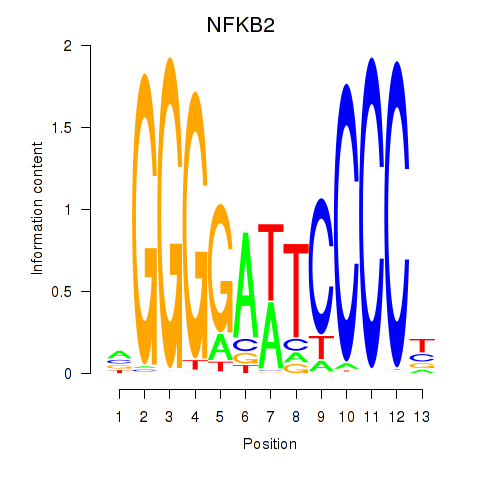
| NFKB2 | 0.026 | 0.097 |
|
|
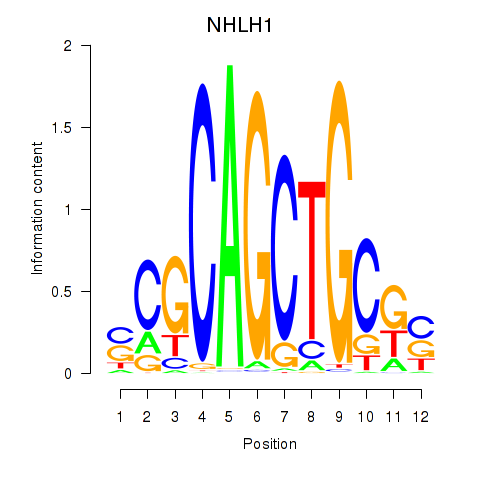
| NHLH1 | 0.038 | 0.153 |
|
|
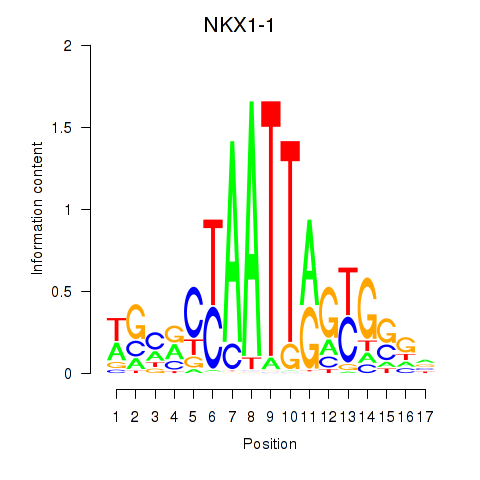
| NKX1-1 | -0.360 | -0.476 |
|
|
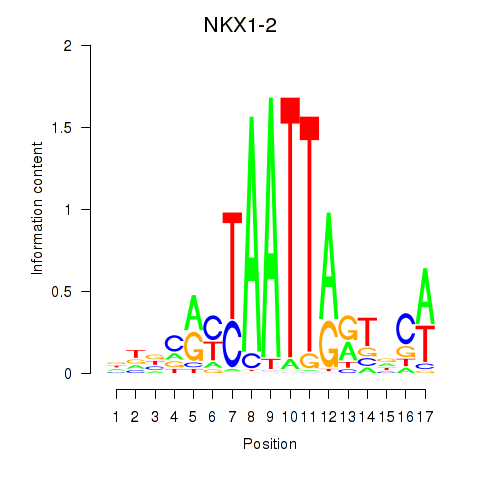
| NKX1-2_RAX | 0.646 | 0.868 |
|
|
| NKX2-1 | -0.222 | -0.416 |
|
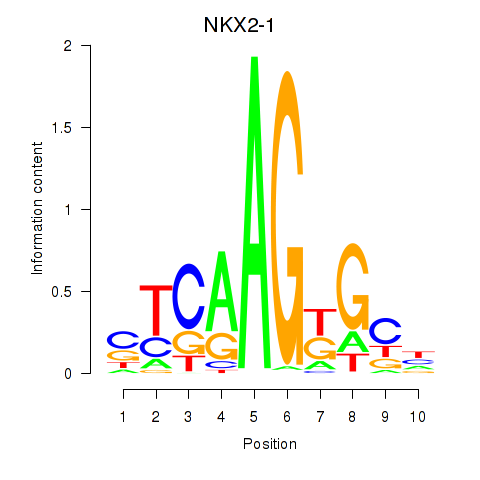
|
| NKX2-2 | 0.319 | 0.922 |
|
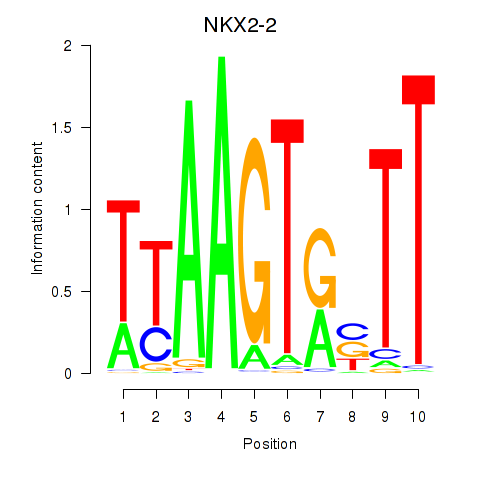
|
| NKX2-3 | 0.076 | 0.224 |
|
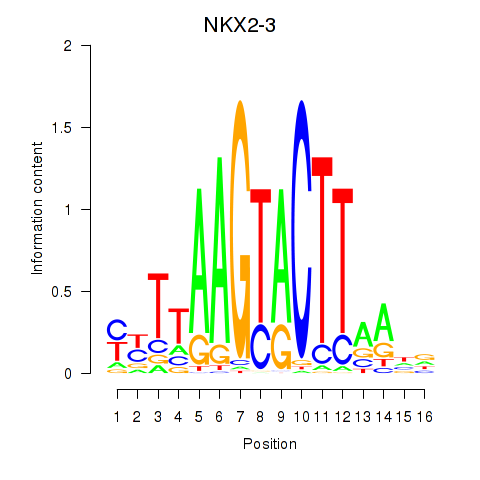
|
| NKX2-4 | -0.031 | -0.095 |
|
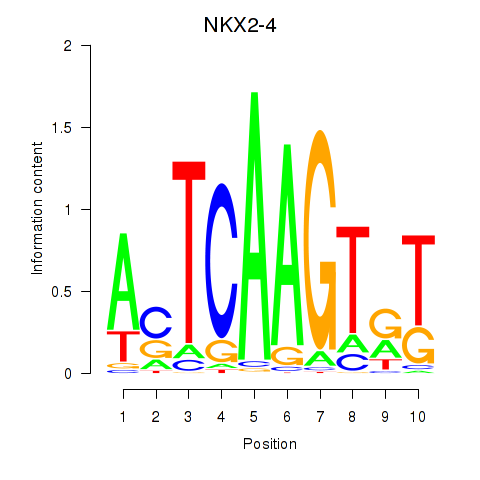
|
| NKX2-5 | -0.120 | -0.235 |
|
|
| NKX2-6 | -0.041 | -0.095 |
|
|
| NKX2-8 | -0.054 | -0.118 |
|
|
| NKX3-1 | 0.116 | 0.291 |
|
|
| NKX3-2 | -0.254 | -0.765 |
|
|
| NKX6-2 | -0.157 | -0.343 |
|
|
| NKX6-3 | -0.285 | -0.217 |
|
|
| NOTO_VSX2_DLX2_DLX6_NKX6-1 | -0.019 | -0.036 |
|
|
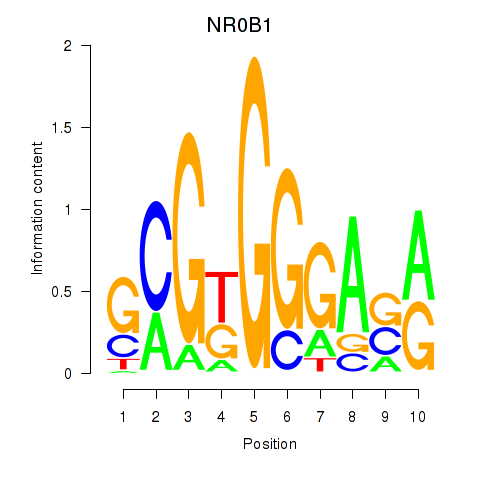
| NR0B1 | 0.089 | 0.246 |
|
|
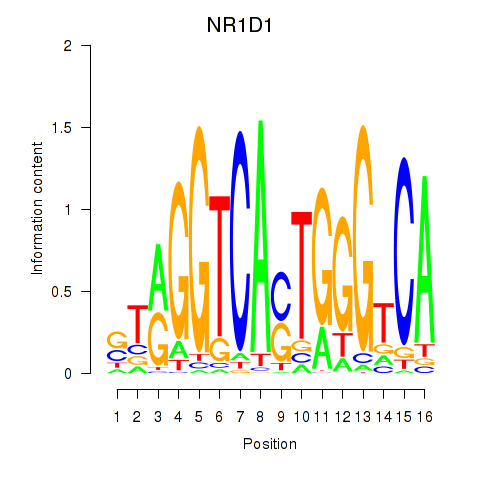
| NR1D1 | -0.203 | -0.327 |
|
|
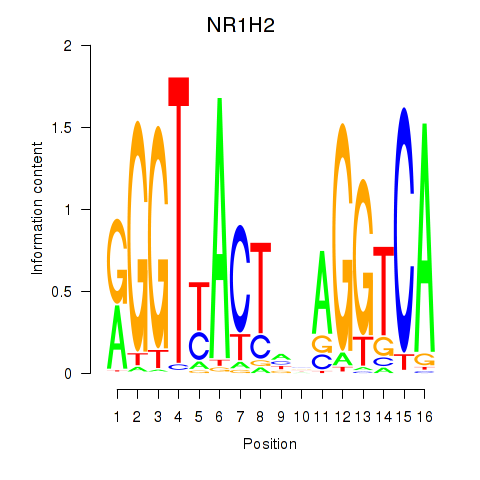
| NR1H2 | -0.078 | -0.181 |
|
|
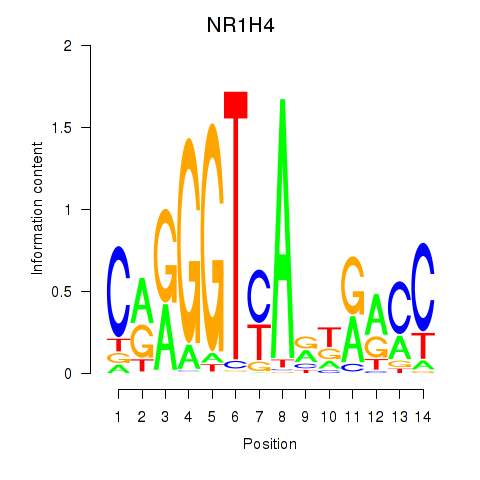
| NR1H4 | -0.463 | -1.046 |
|
|
| NR1I2 | 0.134 | 0.401 |
|
|
| NR1I3 | 0.159 | 0.230 |
|
|
| NR2C1 | 0.338 | 0.597 |
|
|
| NR2E1 | -0.312 | -0.888 |
|
|
| NR2E3 | 0.014 | 0.052 |
|
|
| NR2F1 | -0.185 | -0.466 |
|
|
| NR2F2 | 0.194 | 0.790 |
|
|
| NR3C1 | 0.021 | 0.114 |
|
|
| NR4A1 | -0.320 | -0.584 |
|
|
| NR4A2 | 0.074 | 0.157 |
|
|
| NR4A3 | 0.128 | 0.463 |
|
|
| NR5A2 | -0.120 | -0.513 |
|
|
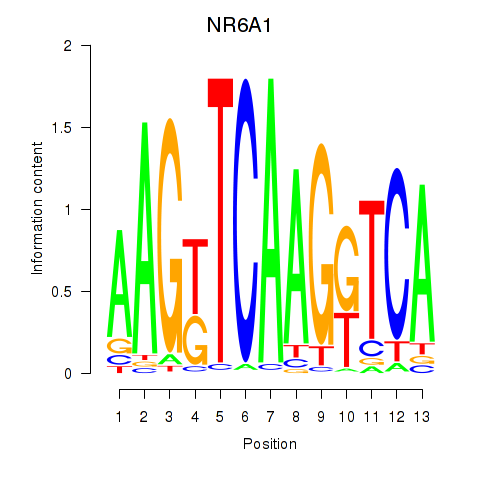
| NR6A1 | -0.234 | -0.718 |
|
|
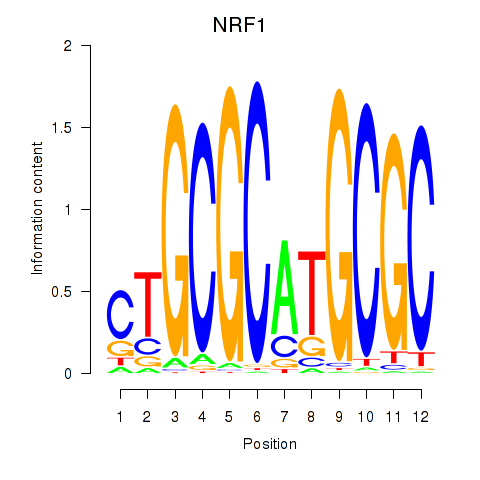
| NRF1 | 0.344 | 4.054 |
|
|
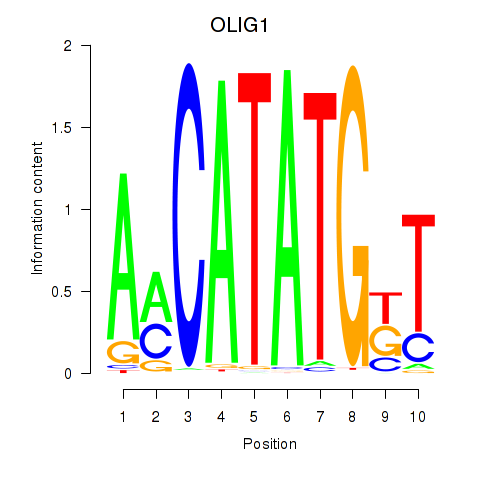
| OLIG1 | -0.218 | -0.434 |
|
|
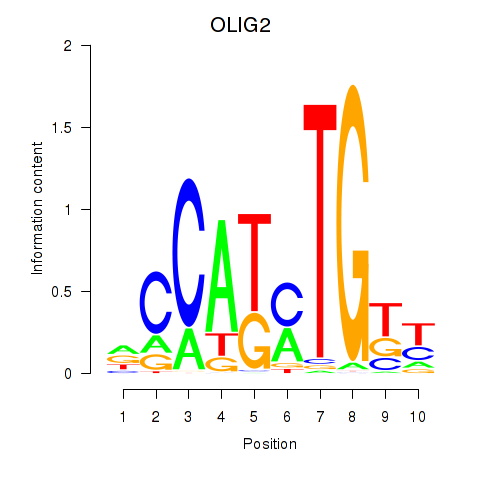
| OLIG2_NEUROD1_ATOH1 | 0.042 | 0.147 |
|
|
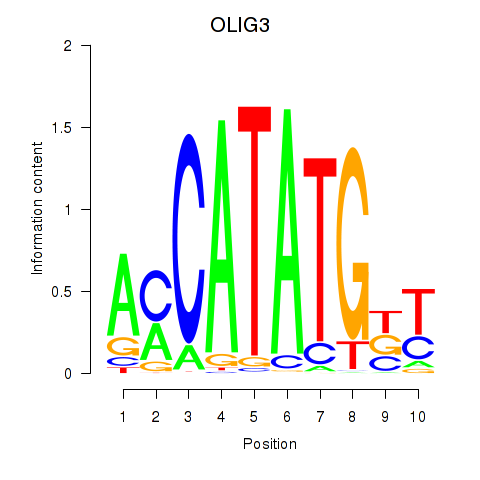
| OLIG3_NEUROD2_NEUROG2 | 0.022 | 0.062 |
|
|
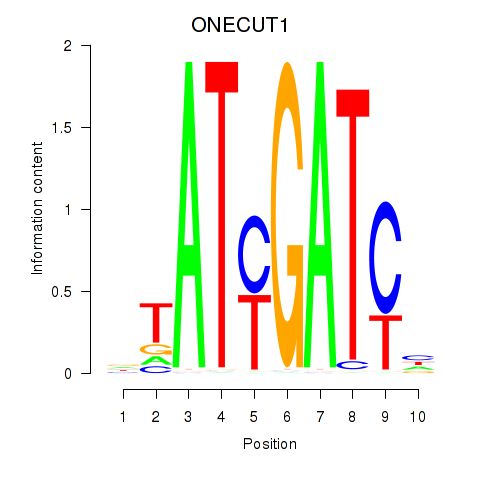
| ONECUT1 | -0.096 | -0.241 |
|
|
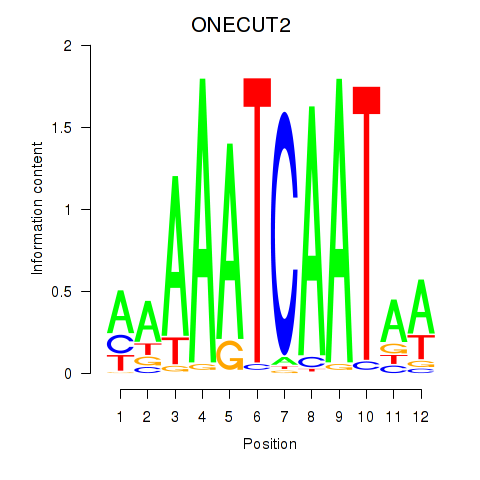
| ONECUT2_ONECUT3 | -0.101 | -0.195 |
|
|
| OSR1_OSR2 | 0.044 | 0.082 |
|

|
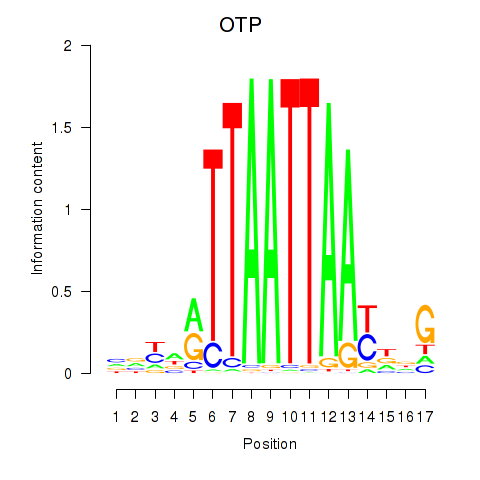
| OTP_PHOX2B_LHX1_LMX1A_LHX5_HOXC4 | 0.160 | 0.296 |
|
|
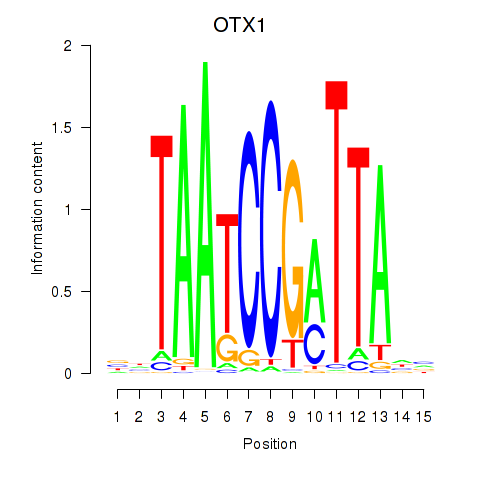
| OTX1 | 0.518 | 1.144 |
|
|
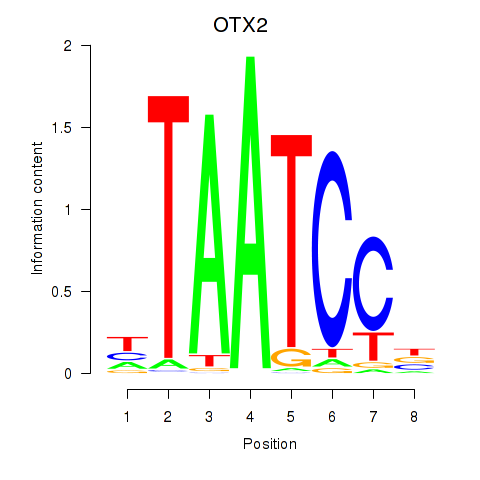
| OTX2_CRX | -0.330 | -0.646 |
|
|
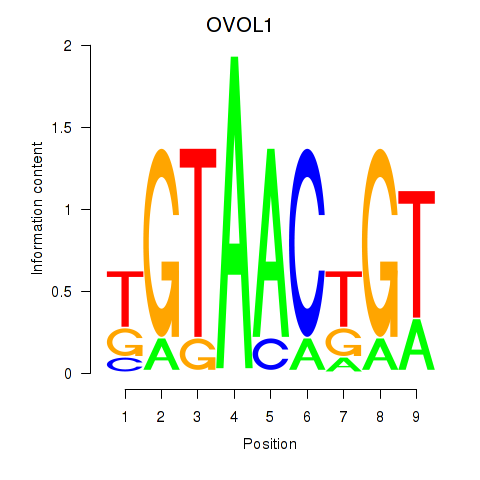
| OVOL1 | 0.807 | 0.514 |
|
|
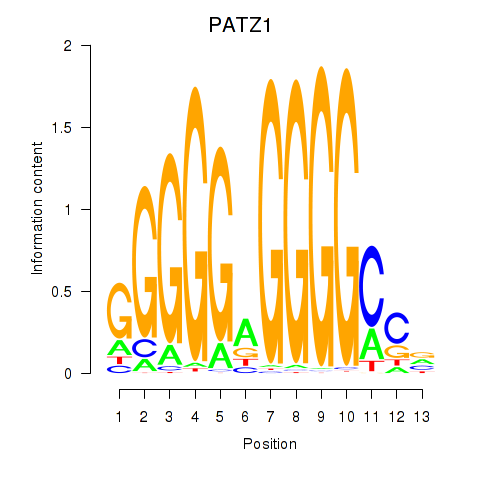
| PATZ1_KLF4 | -0.014 | -0.311 |
|
|
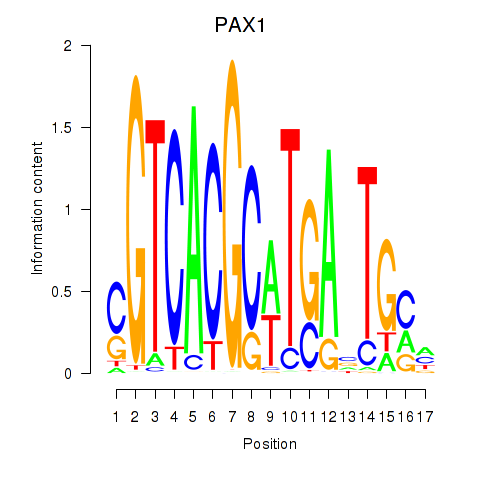
| PAX1_PAX9 | 0.186 | 0.248 |
|
|
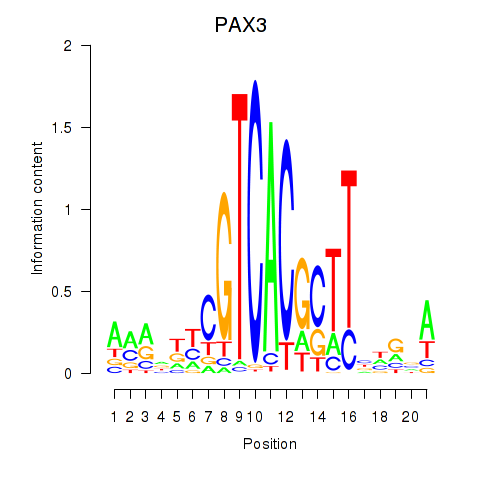
| PAX3 | -0.138 | -0.268 |
|
|
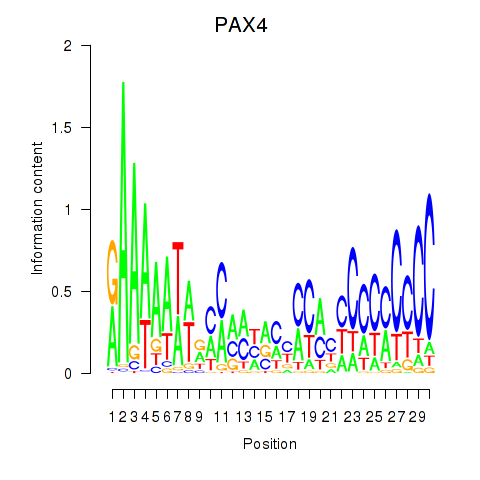
| PAX4 | 0.101 | 0.257 |
|
|
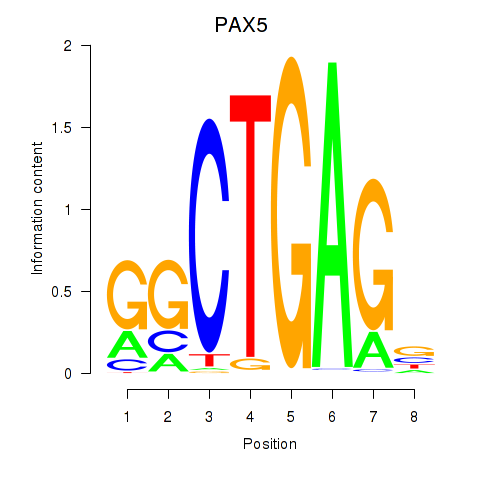
| PAX5 | -0.028 | -0.382 |
|
|
| PAX6 | -0.053 | -0.136 |
|
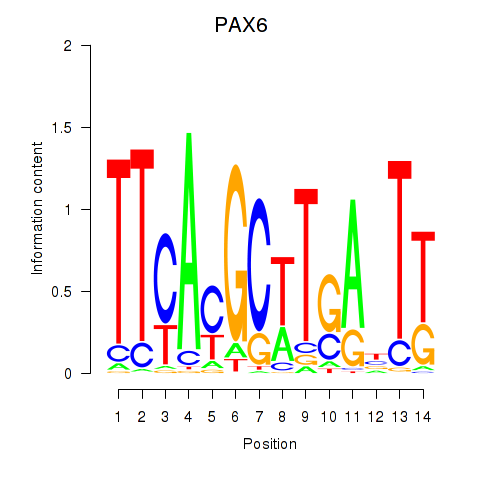
|
| PAX7_NOBOX | -0.155 | -0.398 |
|
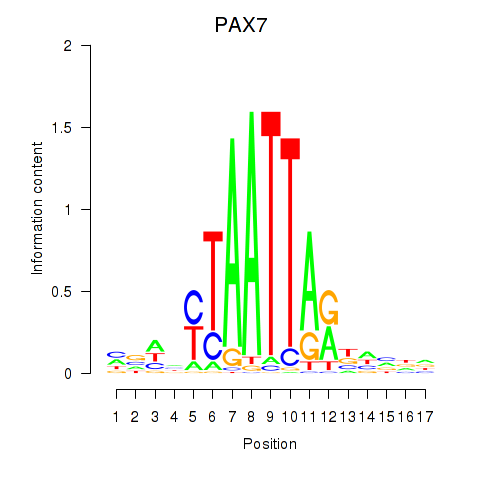
|
| PAX8 | -0.277 | -0.644 |
|
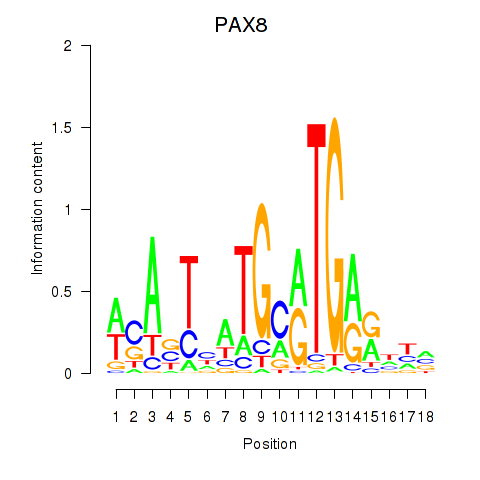
|
| PBX1 | 0.176 | 0.415 |
|
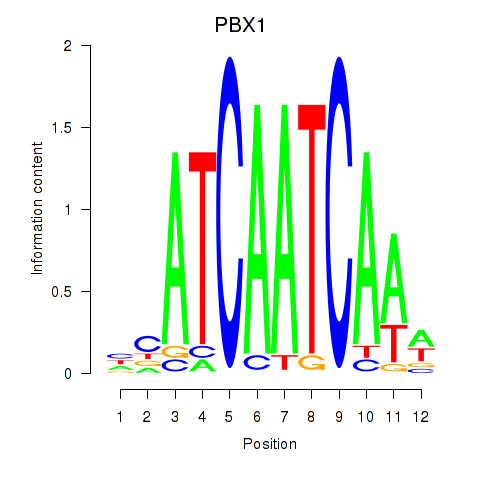
|
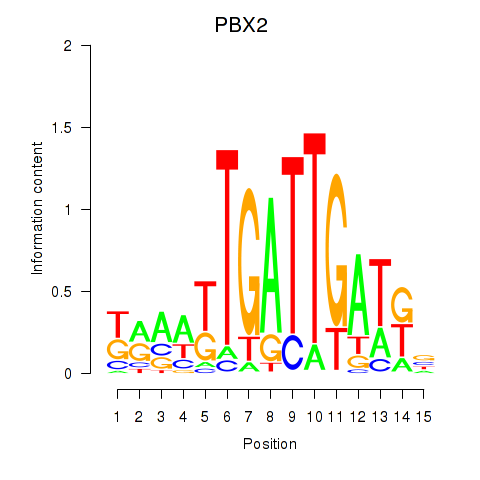
| PBX2 | 0.546 | 0.530 |
|
|
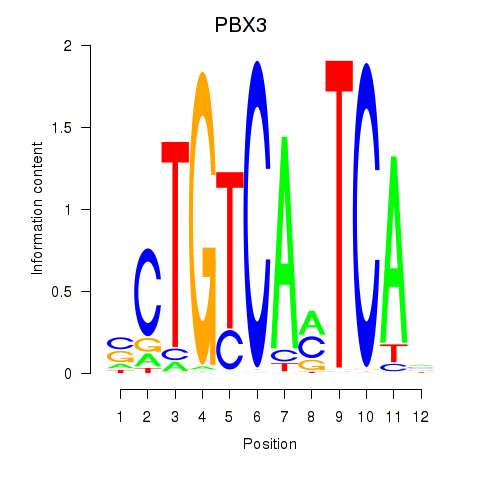
| PBX3 | 0.030 | 0.128 |
|
|
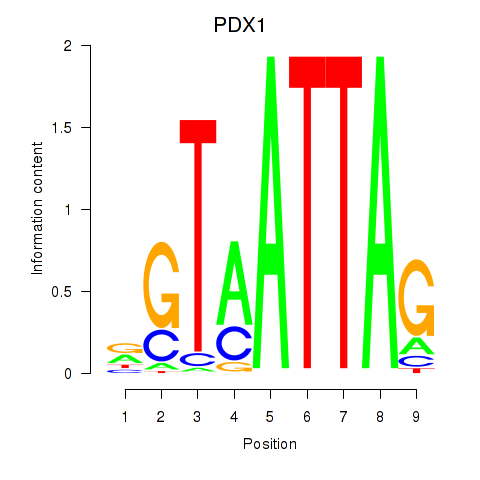
| PDX1 | 0.095 | 0.176 |
|
|
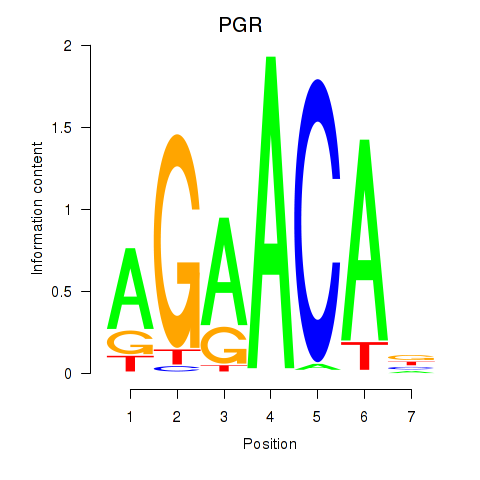
| PGR | -0.082 | -0.225 |
|
|
| PITX1 | 0.109 | 0.994 |
|
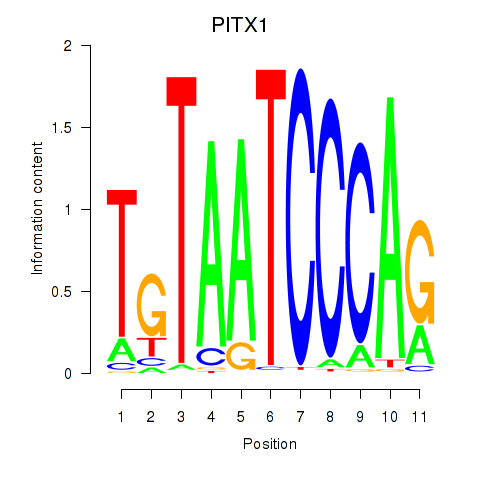
|
| PITX2 | -0.461 | -0.450 |
|
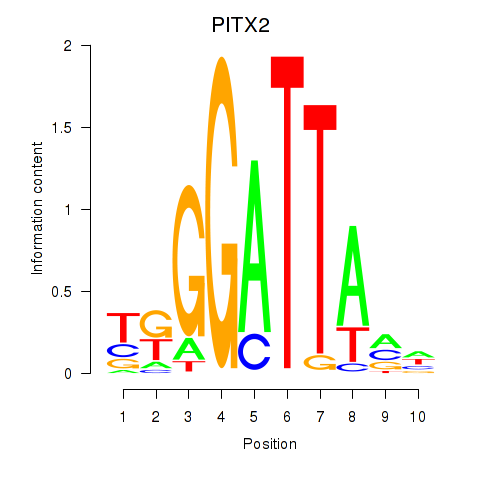
|
| PITX3 | 0.259 | 1.787 |
|
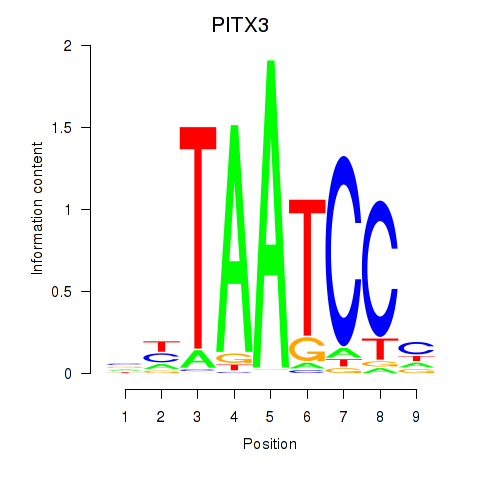
|
| PKNOX1_TGIF2 | -0.145 | -0.377 |
|
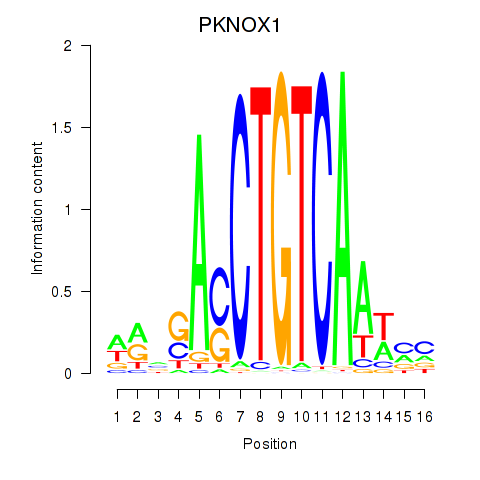
|
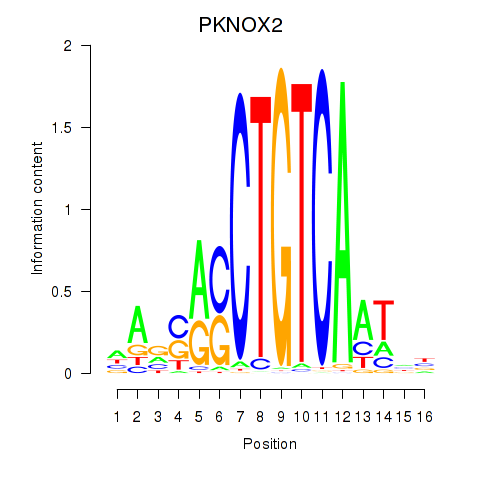
| PKNOX2 | 0.374 | 0.581 |
|
|
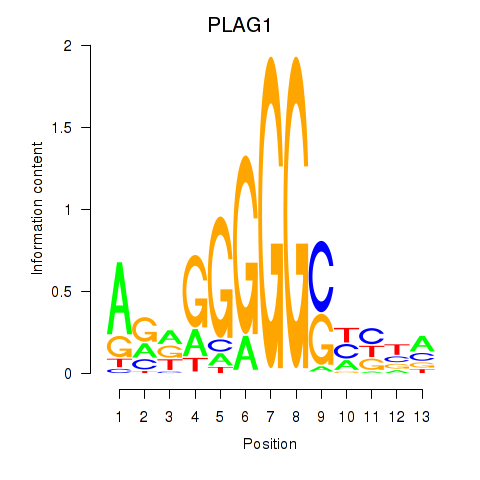
| PLAG1 | -0.390 | -0.737 |
|
|
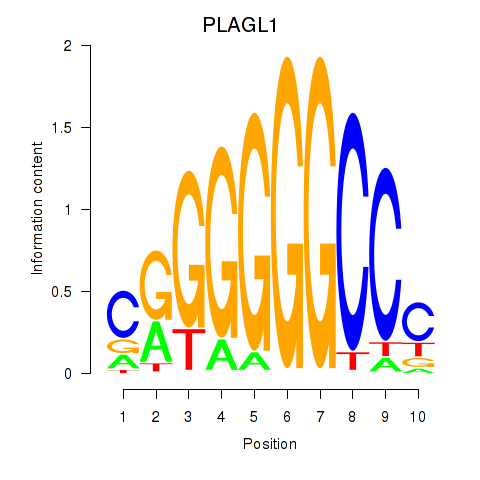
| PLAGL1 | -0.095 | -1.489 |
|
|
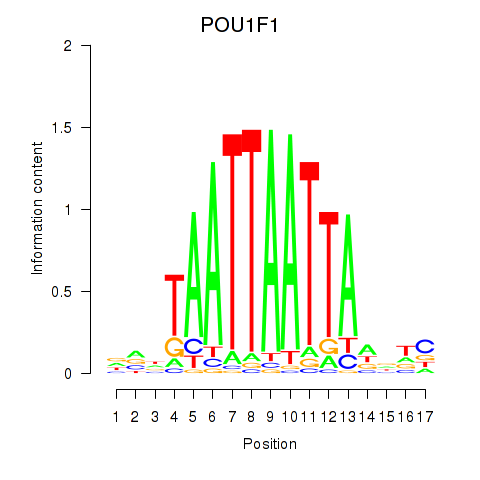
| POU1F1 | 0.202 | 0.435 |
|
|
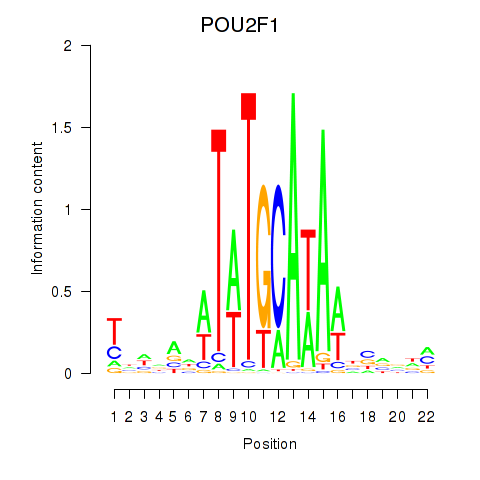
| POU2F1 | 0.186 | 0.398 |
|
|
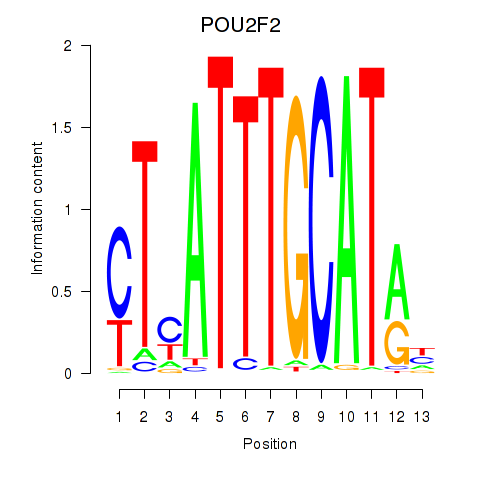
| POU2F2_POU3F1 | -0.205 | -0.652 |
|
|
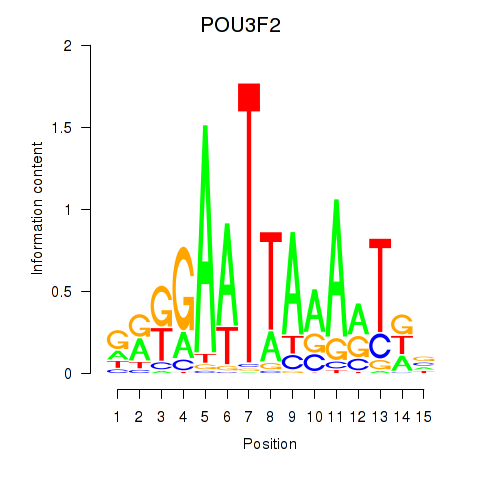
| POU3F2 | 0.118 | 0.941 |
|
|
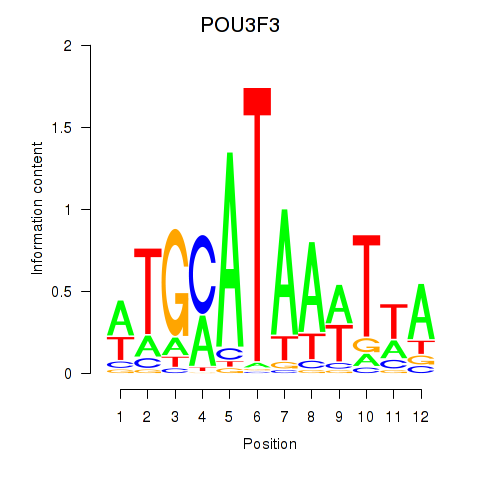
| POU3F3_POU3F4 | -0.256 | -0.714 |
|
|
| POU4F1_POU4F3 | 0.022 | 0.054 |
|
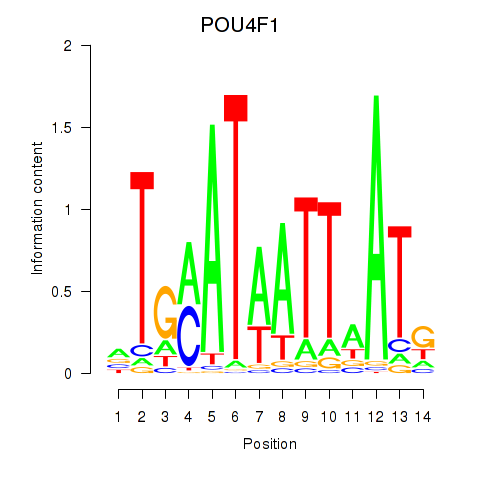
|
| POU4F2 | -0.024 | -0.031 |
|
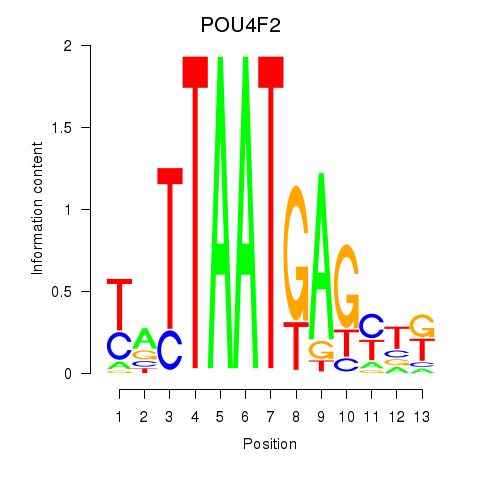
|
| POU5F1_POU2F3 | -0.348 | -1.566 |
|
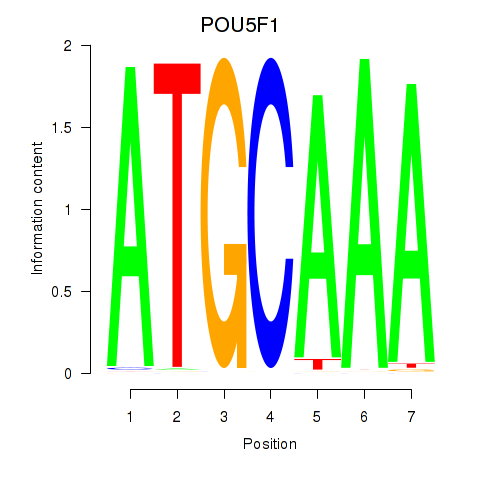
|
| POU6F1 | 0.601 | 0.833 |
|
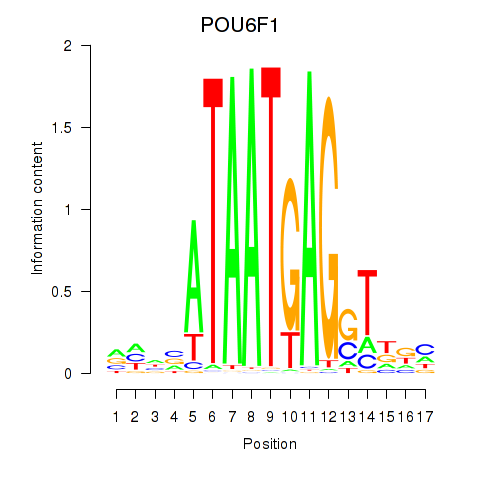
|
| POU6F2 | -0.060 | -0.102 |
|
|
| PPARA | 0.122 | 0.406 |
|
|
| PPARD | -0.457 | -0.701 |
|
|
| PPARG | -0.130 | -0.441 |
|
|
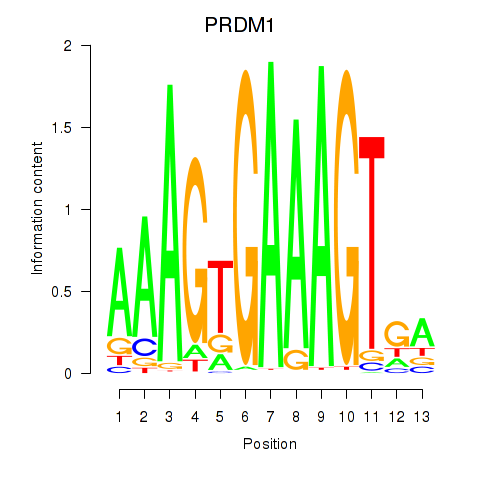
| PRDM1 | -0.240 | -0.570 |
|
|
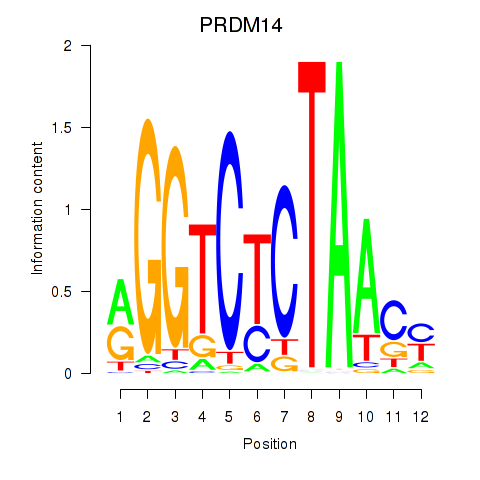
| PRDM14 | -0.016 | -0.019 |
|
|
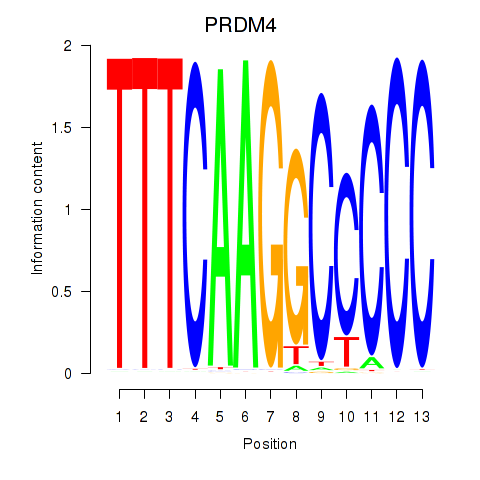
| PRDM4 | -0.079 | -0.102 |
|
|
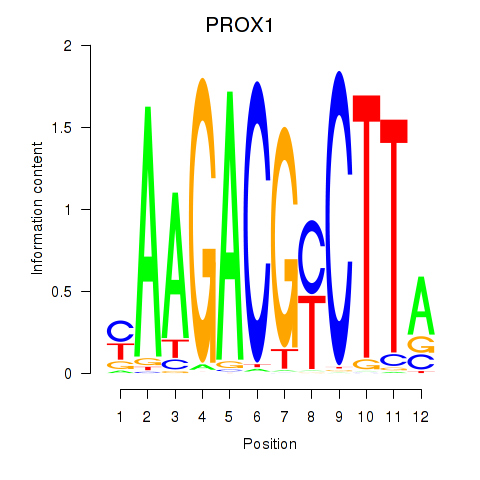
| PROX1 | 0.174 | 0.443 |
|
|
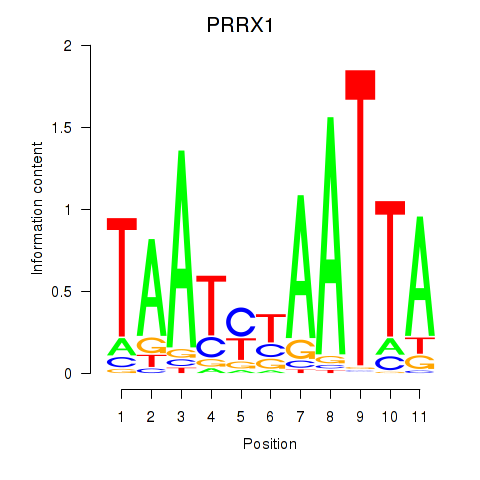
| PRRX1_ALX4_PHOX2A | 0.675 | 1.036 |
|
|
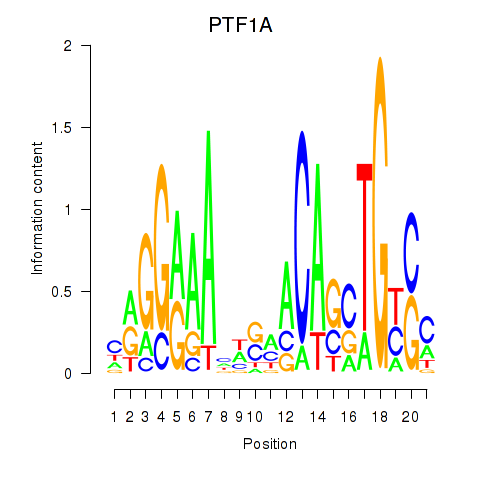
| PTF1A | -0.340 | -0.615 |
|
|
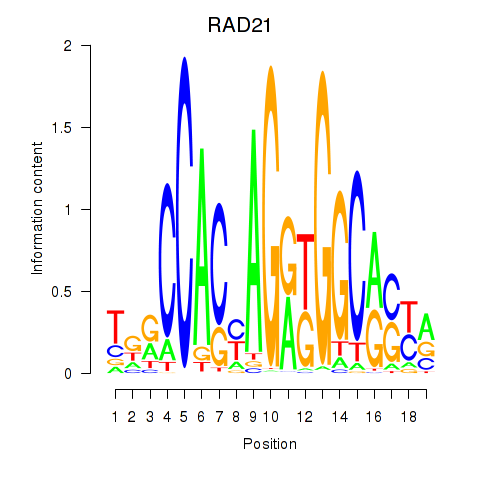
| RAD21_SMC3 | 0.048 | 0.167 |
|
|
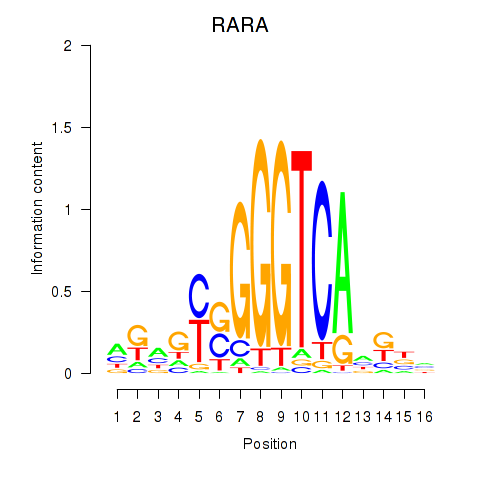
| RARA | -0.154 | -0.478 |
|
|
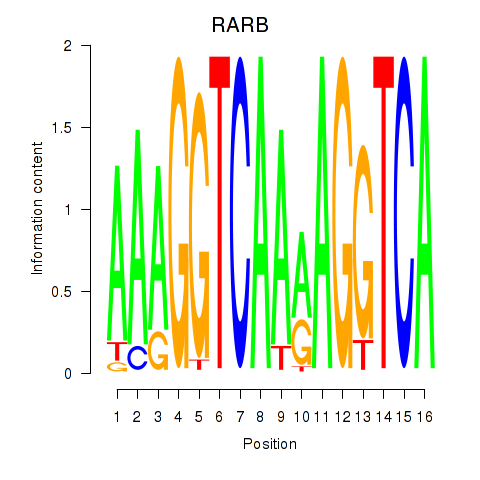
| RARB | -0.341 | -0.837 |
|
|
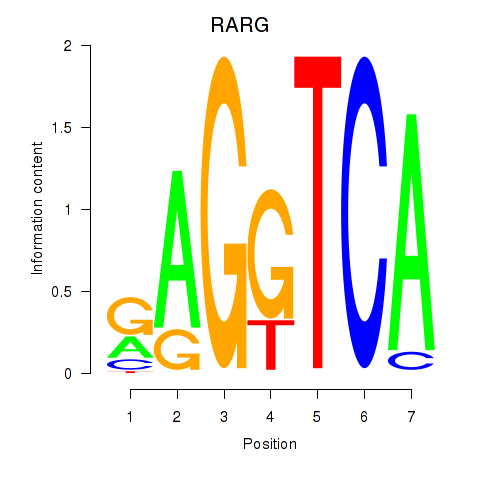
| RARG | 0.055 | 0.296 |
|
|
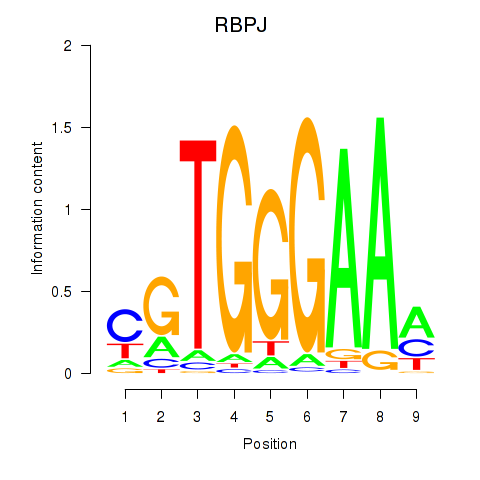
| RBPJ | -0.257 | -0.537 |
|
|
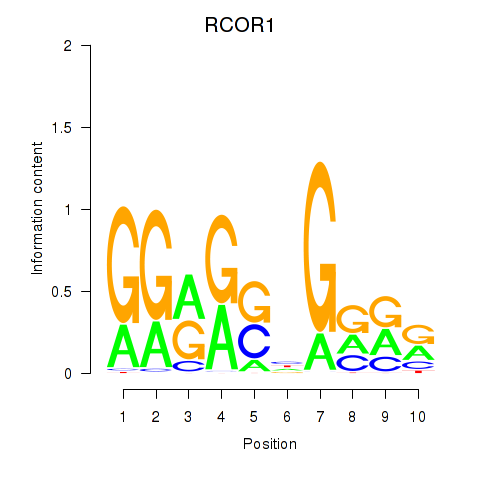
| RCOR1_MTA3 | 0.045 | 1.587 |
|
|
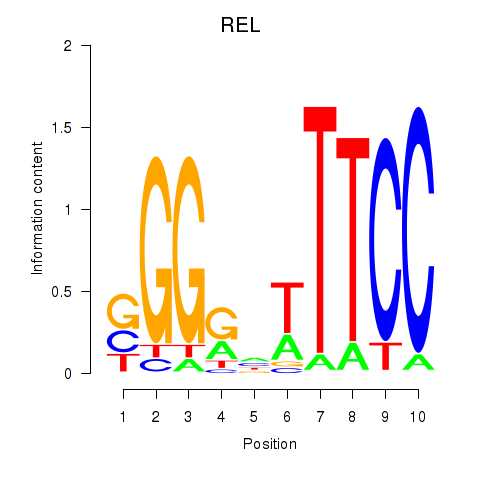
| REL | 0.152 | 0.369 |
|
|
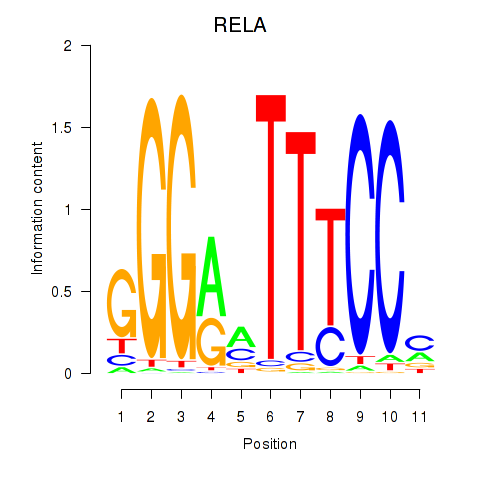
| RELA | 0.076 | 0.174 |
|
|
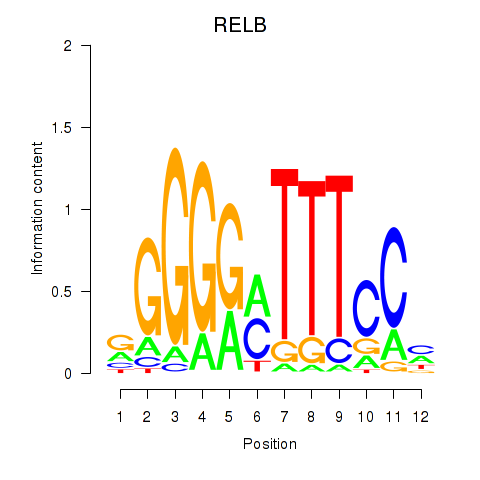
| RELB | -0.174 | -0.470 |
|
|
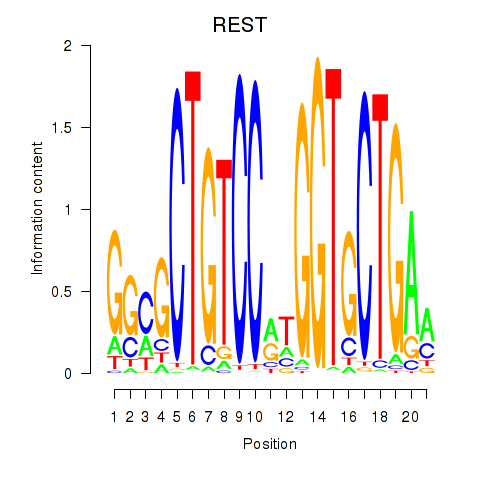
| REST | -1.153 | -2.249 |
|
|
| RFX3_RFX2 | -0.230 | -1.449 |
|
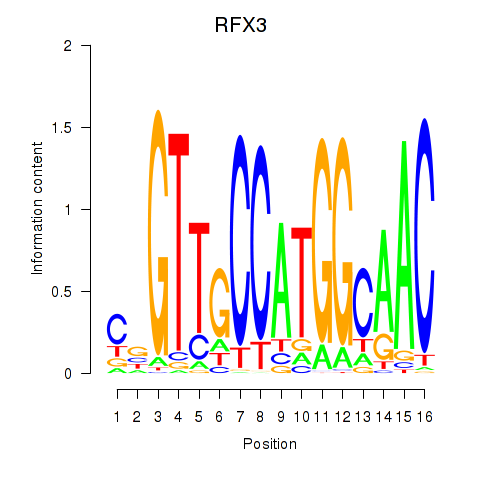
|
| RFX5 | -0.224 | -0.307 |
|
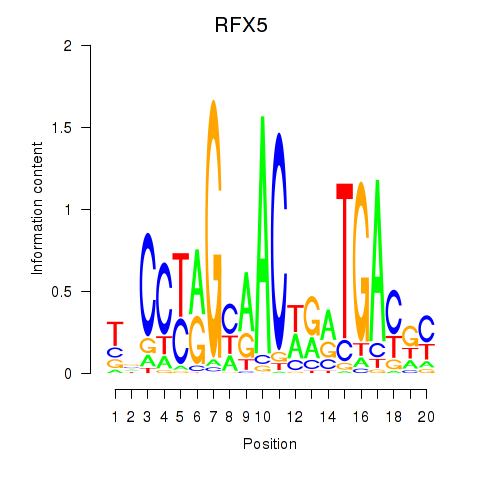
|
| RFX7_RFX4_RFX1 | -0.283 | -1.214 |
|
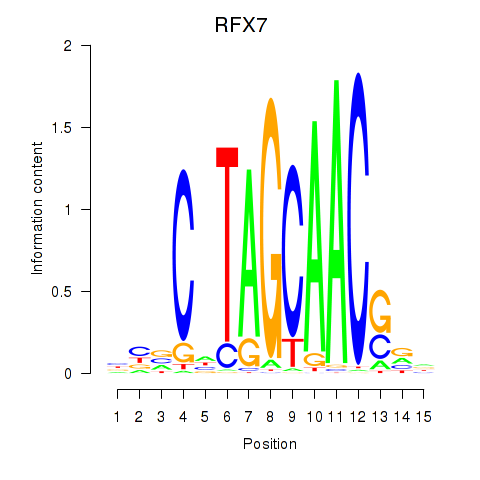
|
| RHOXF1 | -0.248 | -1.368 |
|
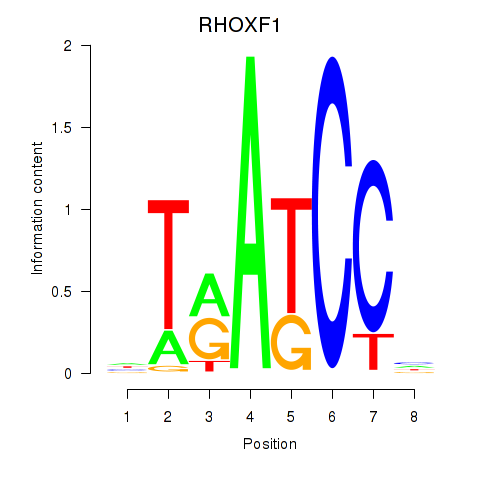
|
| RORA | 0.367 | 0.695 |
|
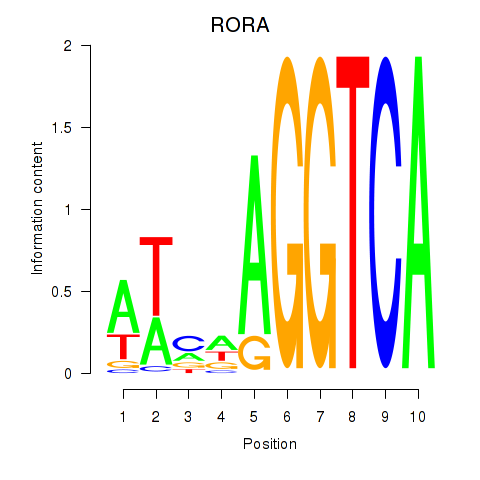
|
| RORC | -0.581 | -0.685 |
|
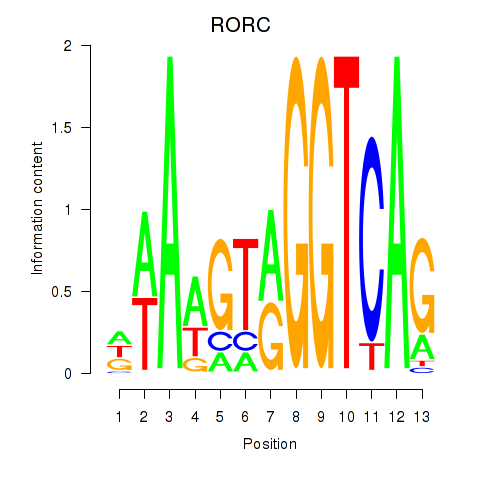
|
| RREB1 | -0.238 | -0.697 |
|
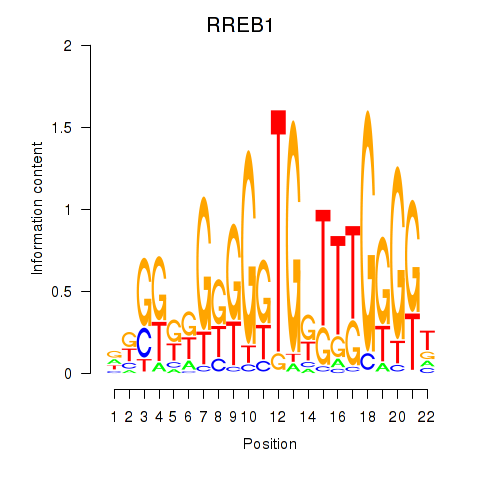
|
| RUNX1_RUNX2 | -0.188 | -1.034 |
|
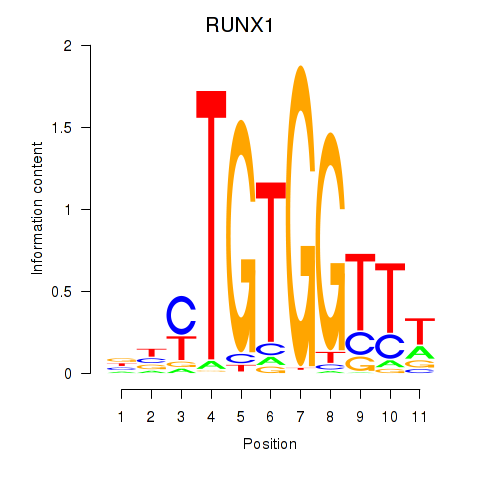
|
| RUNX3_BCL11A | -0.145 | -0.427 |
|
|
| RXRA_NR2F6_NR2C2 | 0.129 | 0.424 |
|
|
| RXRG | -0.097 | -0.447 |
|
|
| SCRT1_SCRT2 | 0.152 | 0.402 |
|
|
| SHOX | -0.225 | -0.432 |
|
|
| SHOX2_HOXC5 | 0.031 | 0.128 |
|
|
| SIN3A_CHD1 | -0.021 | -0.302 |
|
|
| SIX1_SIX3_SIX2 | -0.116 | -0.302 |
|
|
| SIX4 | -0.318 | -0.465 |
|
|
| SIX5_SMARCC2_HCFC1 | 0.149 | 1.915 |
|
|
| SIX6 | 0.077 | 0.139 |
|
|
| SMAD1 | -0.192 | -2.559 |
|
|
| SMAD2 | -0.460 | -1.020 |
|

|
| SMAD3 | -0.417 | -0.808 |
|
|
| SMAD4 | -0.019 | -0.111 |
|
|
| SOX1 | -0.254 | -0.440 |
|
|
| SOX10_SOX15 | 0.372 | 0.867 |
|
|
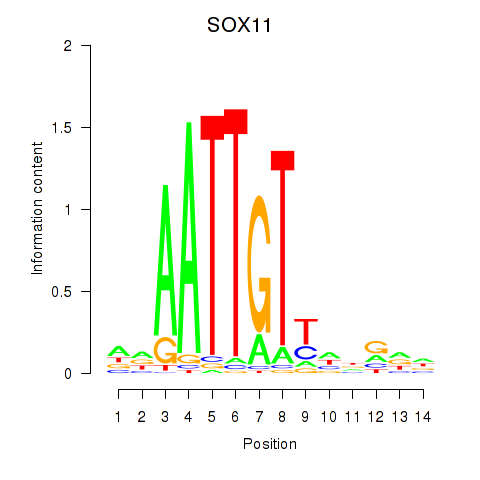
| SOX11 | -0.222 | -0.402 |
|
|
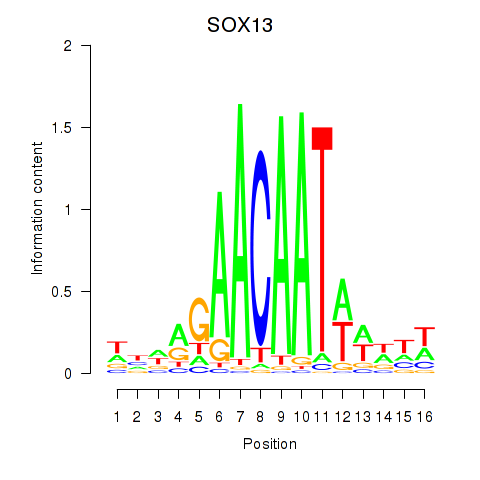
| SOX13_SOX12 | -0.226 | -0.964 |
|
|
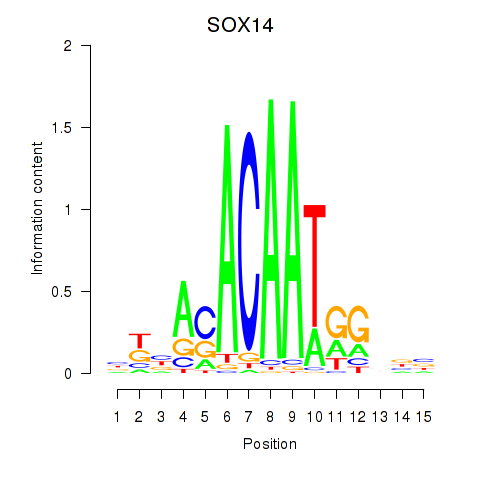
| SOX14 | 0.053 | 0.118 |
|
|
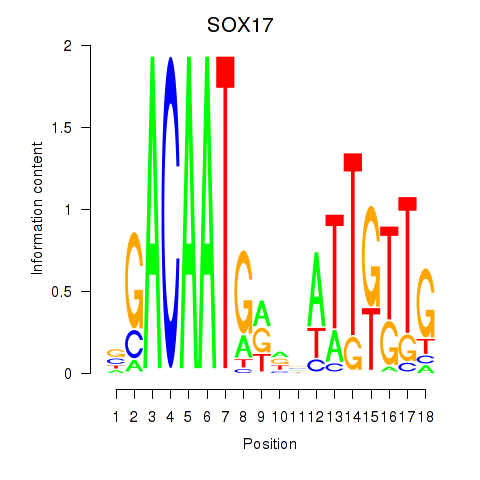
| SOX17 | 0.018 | 0.055 |
|
|
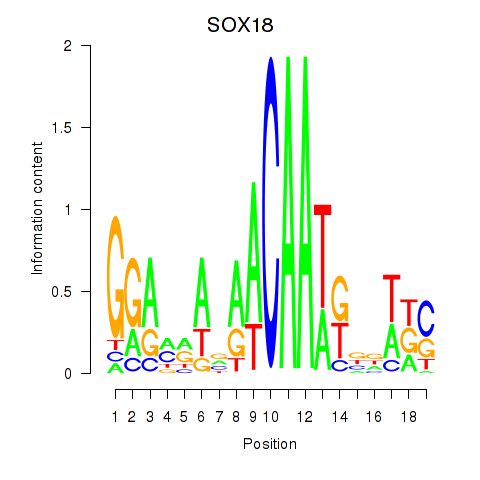
| SOX18 | -0.070 | -0.155 |
|
|
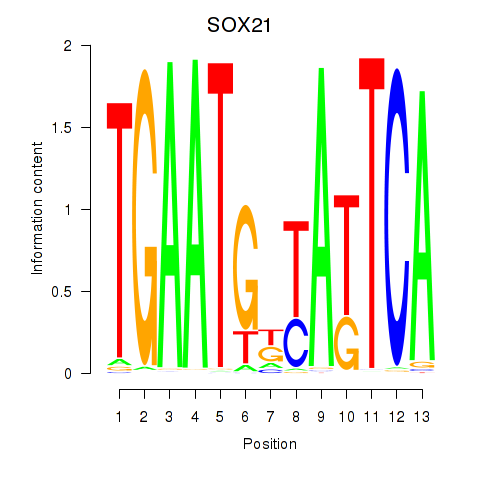
| SOX21 | 0.101 | 0.136 |
|
|
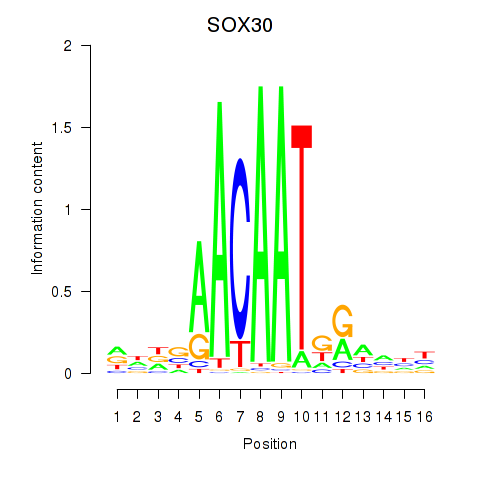
| SOX30 | 0.125 | 0.181 |
|
|
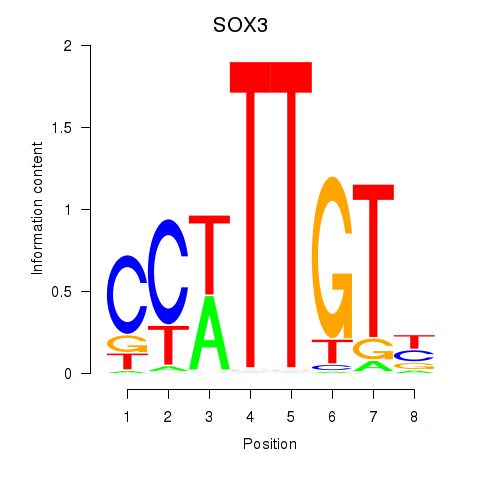
| SOX3_SOX2 | 0.026 | 0.137 |
|
|
| SOX4 | -0.208 | -0.457 |
|
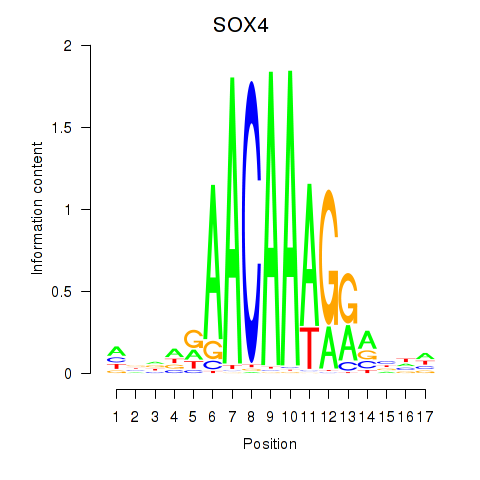
|
| SOX5 | 0.018 | 0.043 |
|
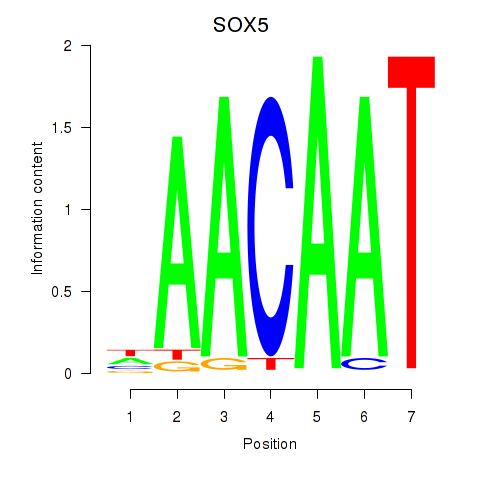
|
| SOX6 | 0.105 | 0.237 |
|
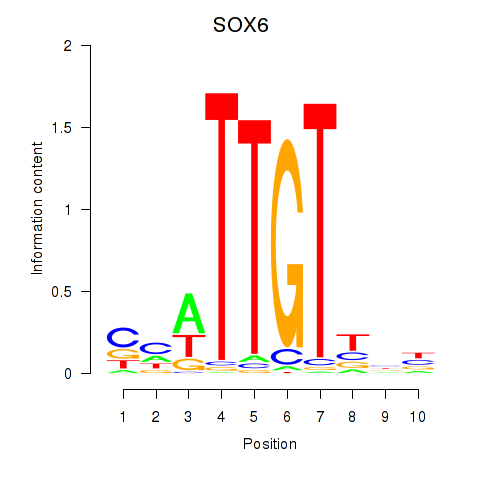
|
| SOX7 | 0.065 | 0.081 |
|
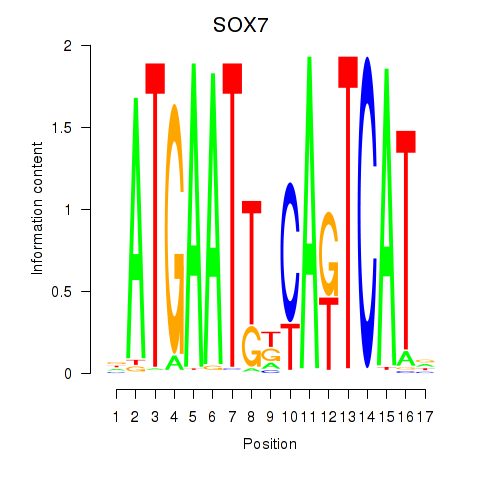
|
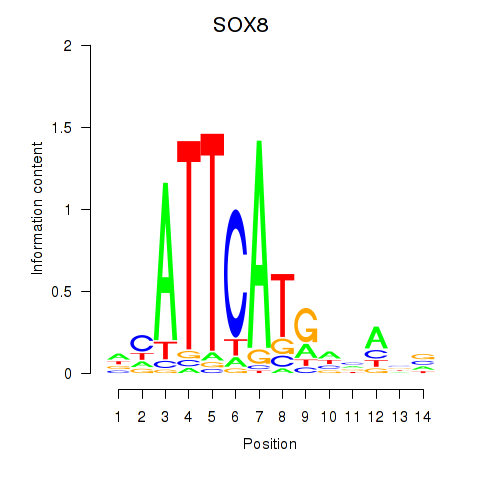
| SOX8 | -0.050 | -0.125 |
|
|
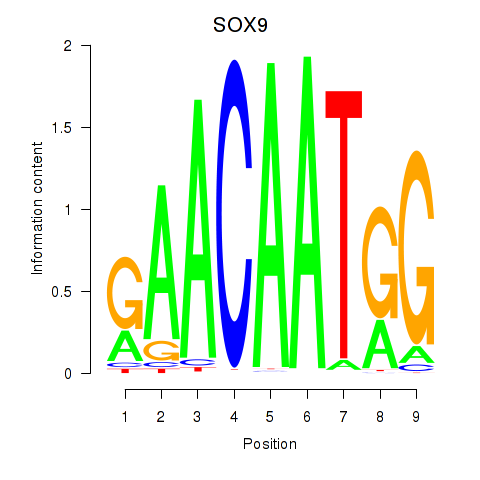
| SOX9 | 0.174 | 0.618 |
|
|
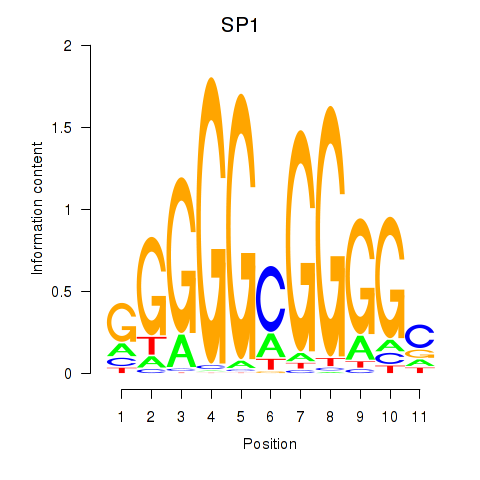
| SP1 | 0.051 | 0.807 |
|
|
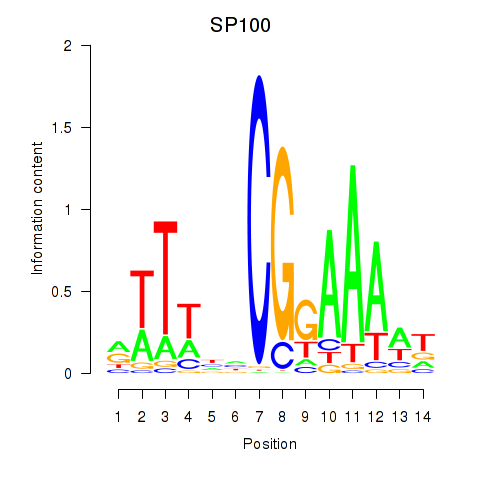
| SP100 | 0.628 | 2.370 |
|
|
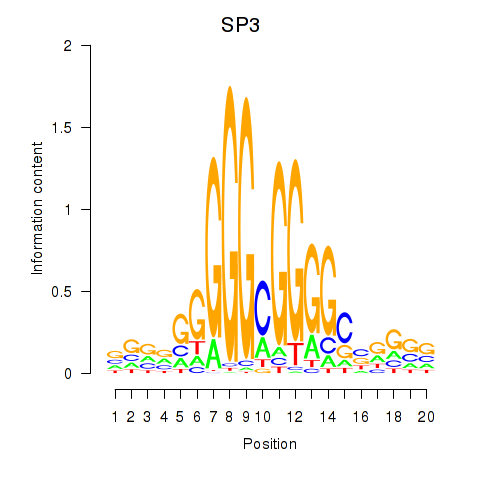
| SP3 | 0.061 | 1.414 |
|
|
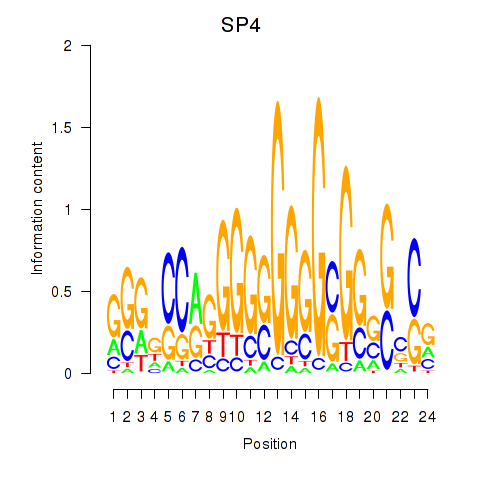
| SP4_PML | -0.040 | -0.964 |
|
|
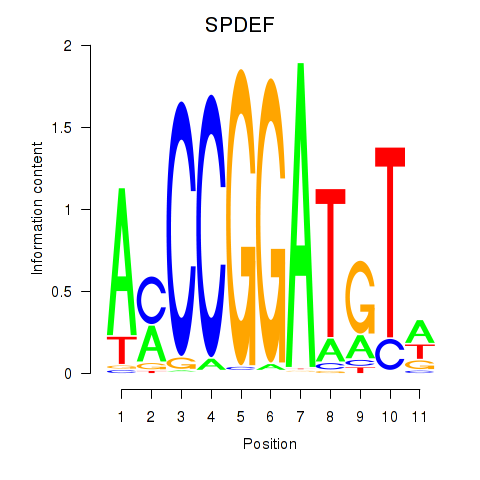
| SPDEF | 0.393 | 1.021 |
|
|
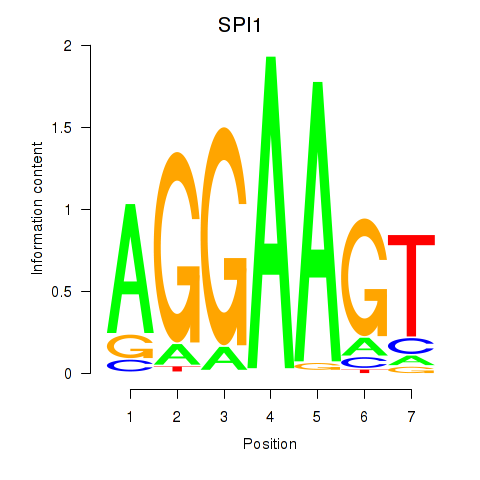
| SPI1 | 0.076 | 0.360 |
|
|
| SPIB | 0.224 | 0.772 |
|
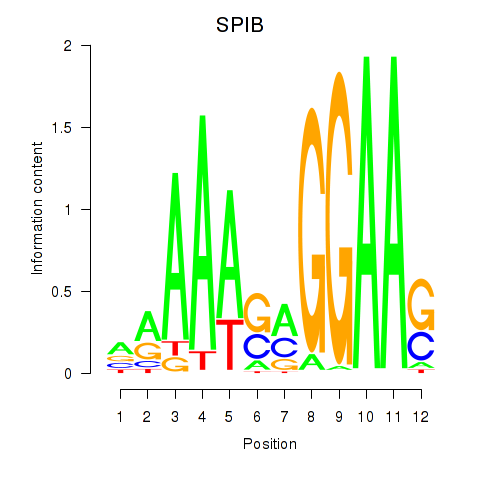
|
| SPIC | -0.098 | -0.469 |
|
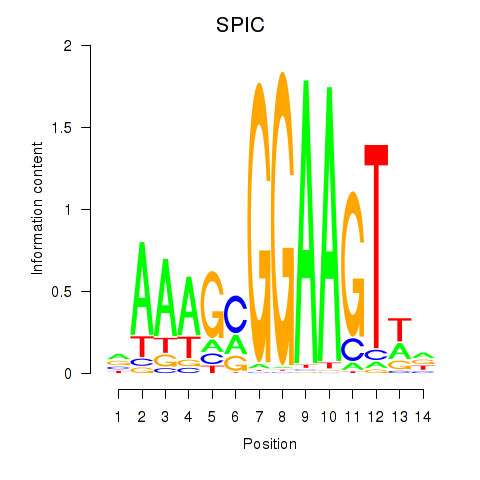
|
| SREBF1_TFE3 | -0.127 | -0.497 |
|
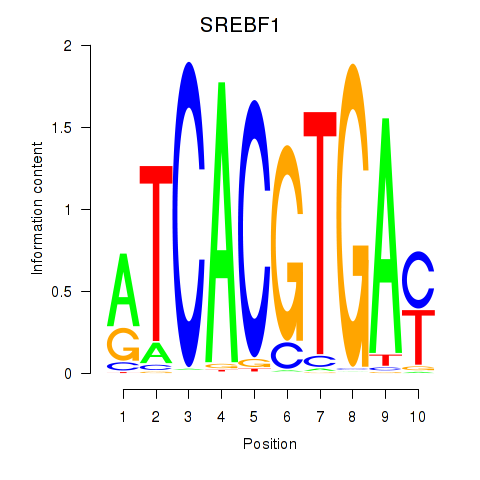
|
| SREBF2 | 0.723 | 1.012 |
|
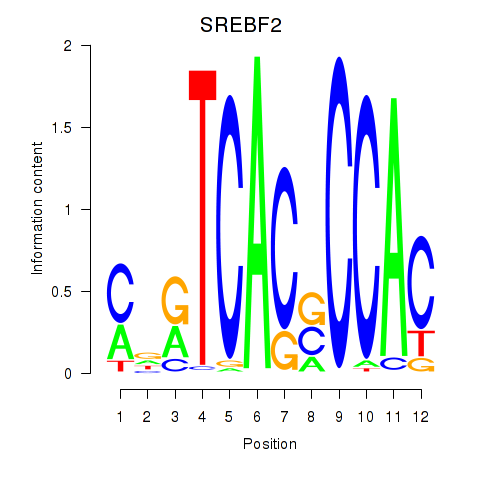
|
| SRF | 0.399 | 0.948 |
|
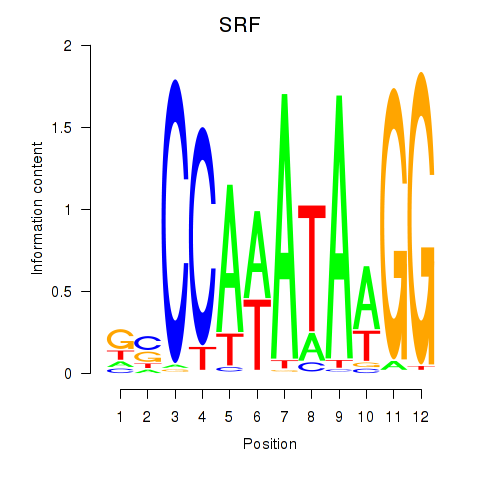
|
| SRY | -0.260 | -0.608 |
|
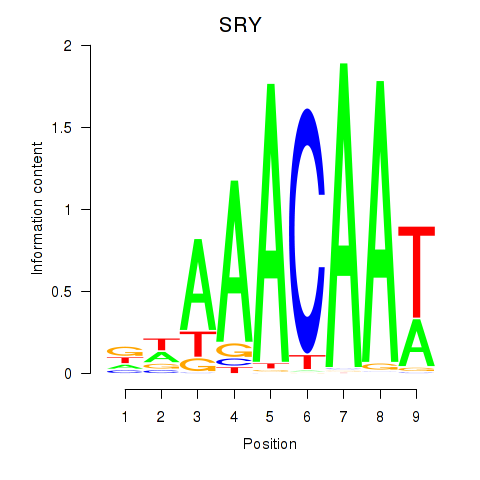
|
| STAT1_STAT3_BCL6 | -0.032 | -0.111 |
|
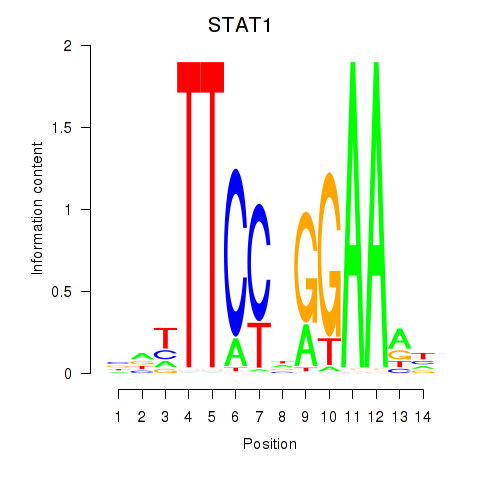
|
| STAT4 | 0.125 | 0.253 |
|
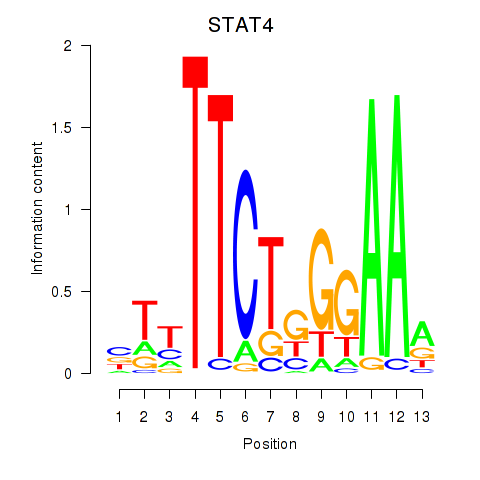
|
| STAT5A | -0.213 | -1.162 |
|
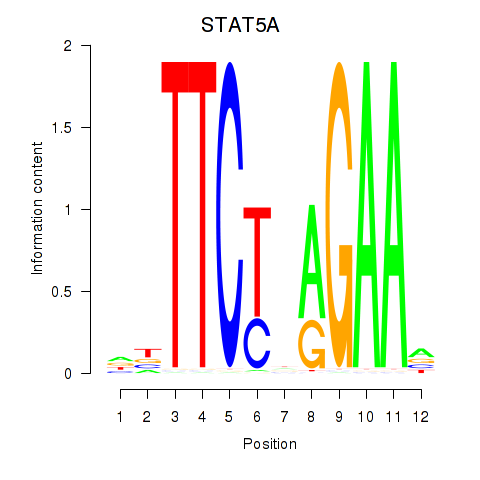
|
| STAT5B | -0.260 | -0.683 |
|
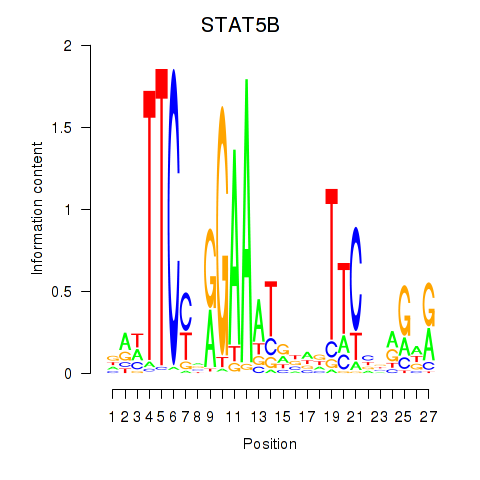
|
| STAT6 | -0.230 | -0.489 |
|
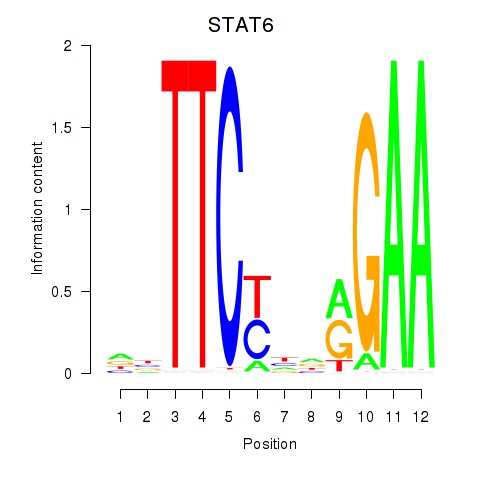
|
| T | 0.135 | 0.258 |
|
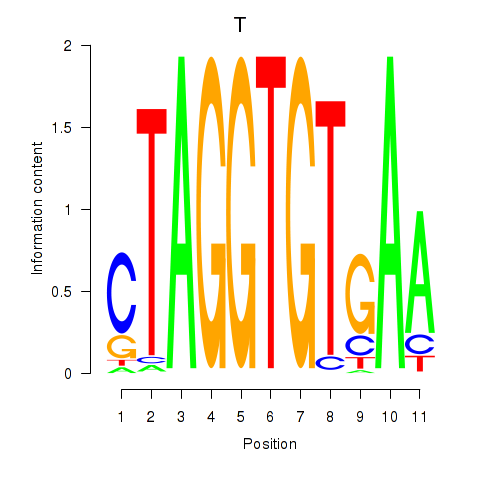
|
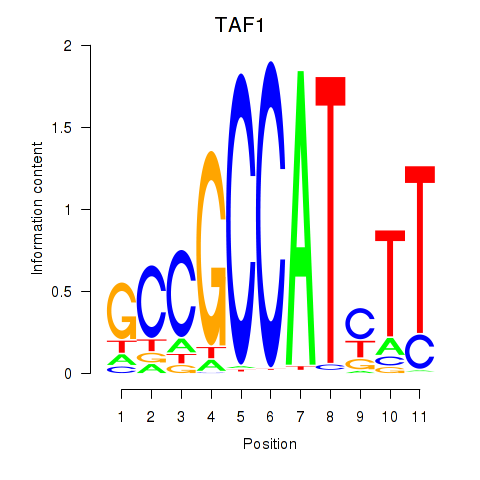
| TAF1 | 0.318 | 1.871 |
|
|
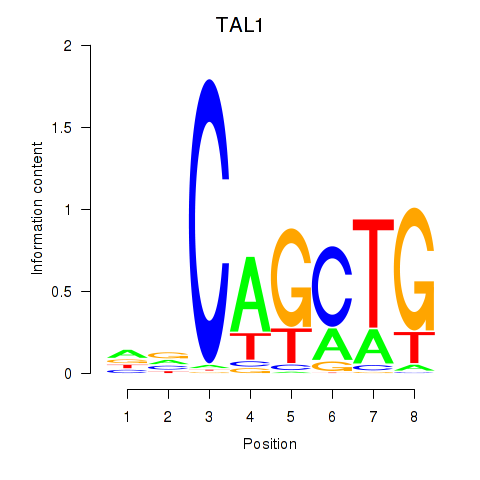
| TAL1 | -0.100 | -1.201 |
|
|
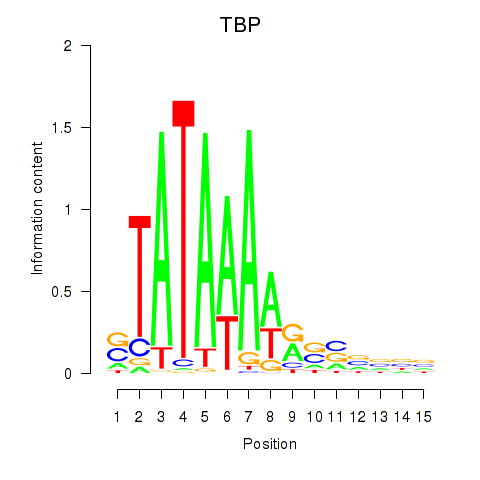
| TBP | 0.126 | 0.488 |
|
|
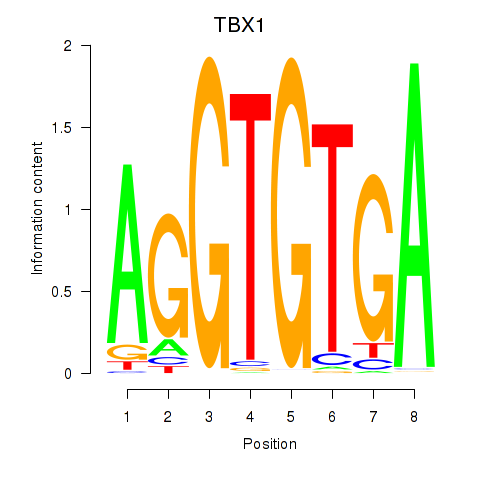
| TBX1 | -0.007 | -0.046 |
|
|
| TBX15_MGA | 0.015 | 0.043 |
|
|
| TBX19 | -0.062 | -0.134 |
|
|
| TBX2 | 0.105 | 0.283 |
|
|
| TBX20 | 0.246 | 0.366 |
|
|
| TBX21_TBR1 | -0.141 | -0.361 |
|
|
| TBX3 | -0.024 | -0.059 |
|
|
| TBX4 | 0.048 | 0.069 |
|
|
| TBX5 | -0.184 | -0.559 |
|
|
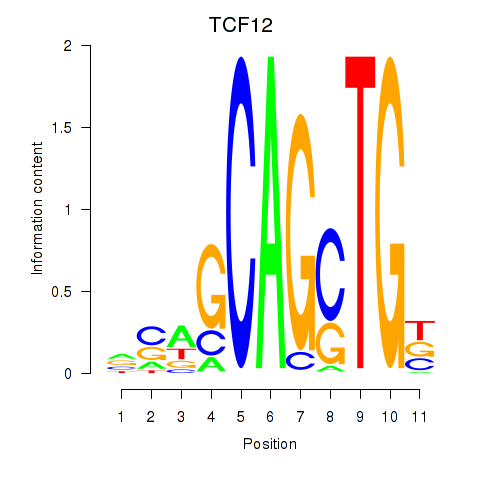
| TCF12_ASCL2 | -0.151 | -0.892 |
|
|
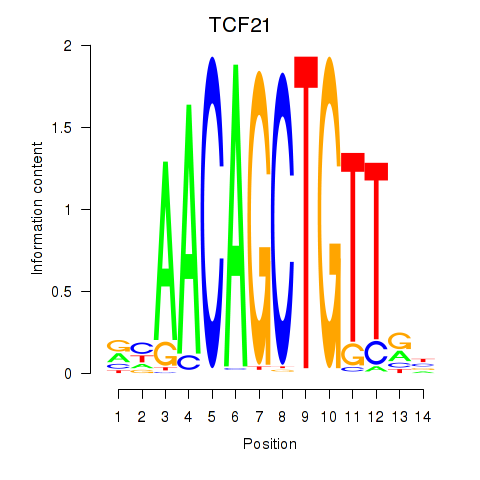
| TCF21 | 0.028 | 0.089 |
|
|
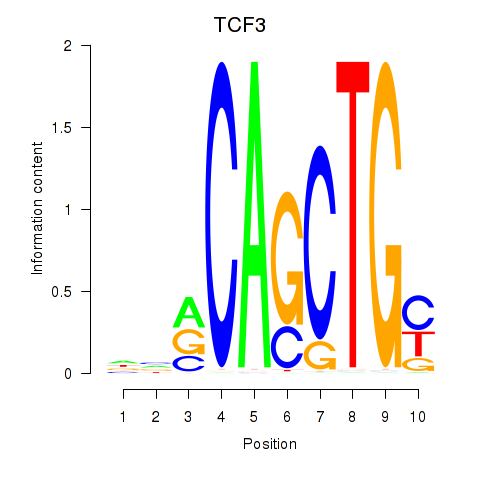
| TCF3_MYOG | 0.157 | 0.610 |
|
|
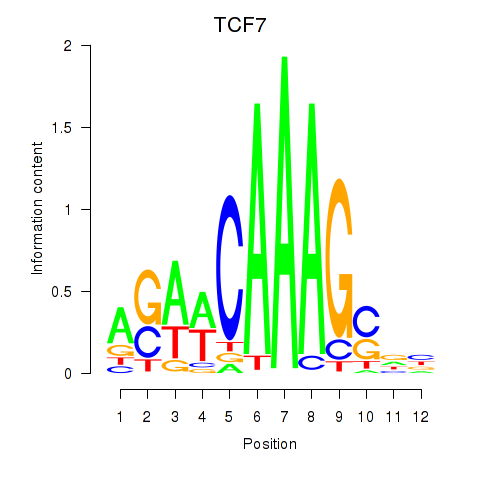
| TCF7 | 0.221 | 0.618 |
|
|
| TCF7L1 | 0.011 | 0.023 |
|
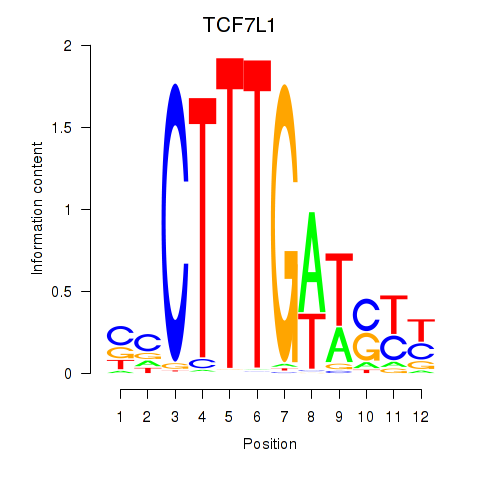
|
| TCF7L2 | 0.128 | 0.252 |
|
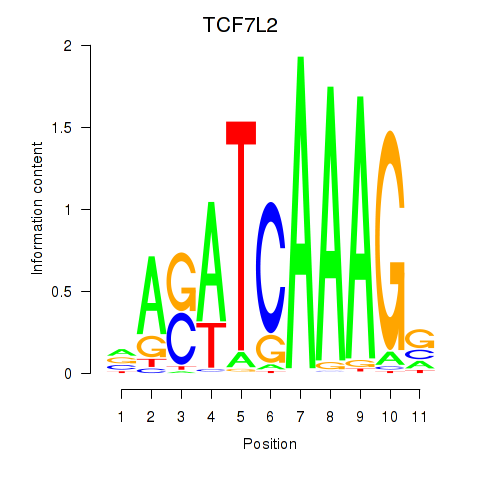
|
| TEAD3_TEAD1 | 0.079 | 0.304 |
|
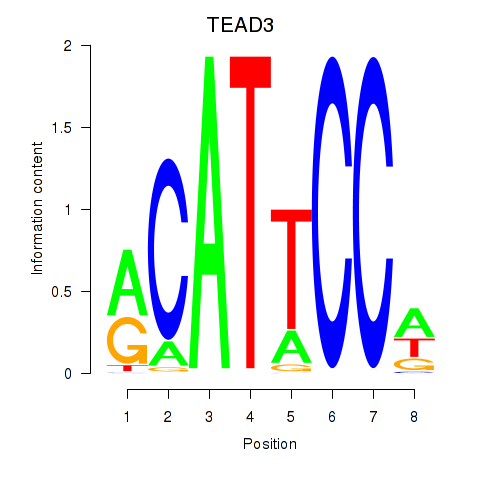
|
| TEAD4 | -0.365 | -0.893 |
|
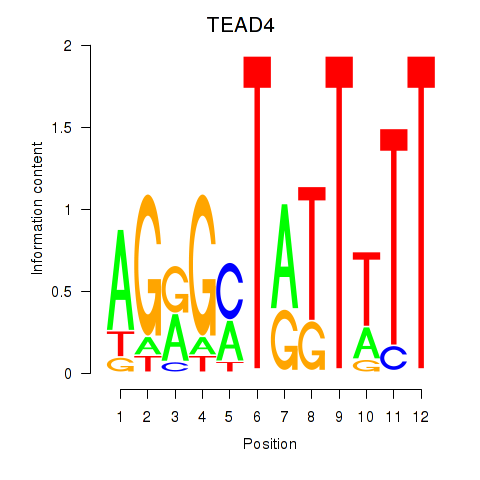
|
| TFAP2B | -0.170 | -0.647 |
|
|
| TFAP2C | -0.144 | -1.326 |
|
|
| TFAP2E | -0.329 | -0.550 |
|
|
| TFAP4_MSC | 0.056 | 0.183 |
|
|
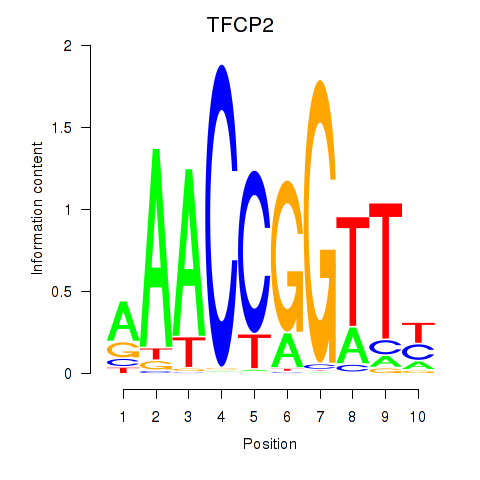
| TFCP2 | 0.258 | 0.746 |
|
|
| TFCP2L1 | 0.005 | 0.012 |
|

|
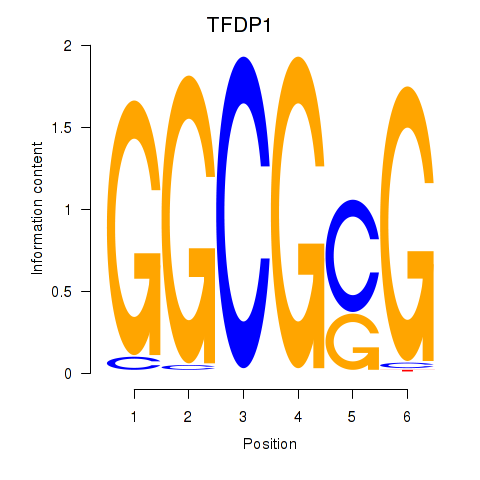
| TFDP1 | 0.043 | 0.381 |
|
|
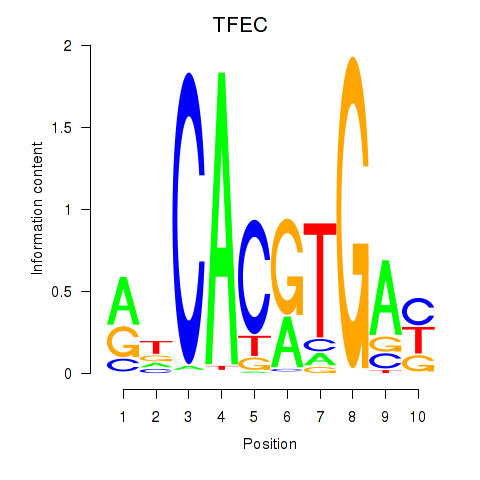
| TFEC_MITF_ARNTL_BHLHE41 | 0.064 | 0.233 |
|
|
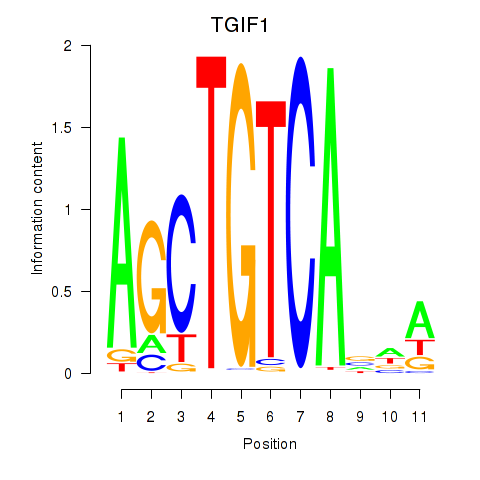
| TGIF1 | -0.199 | -0.695 |
|
|
| THRA_RXRB | 0.023 | 0.061 |
|
|
| THRB | 0.465 | 0.526 |
|
|
| TLX1_NFIC | -0.015 | -0.053 |
|
|
| TLX2 | 0.035 | 0.134 |
|
|
| TP53 | 0.062 | 0.175 |
|
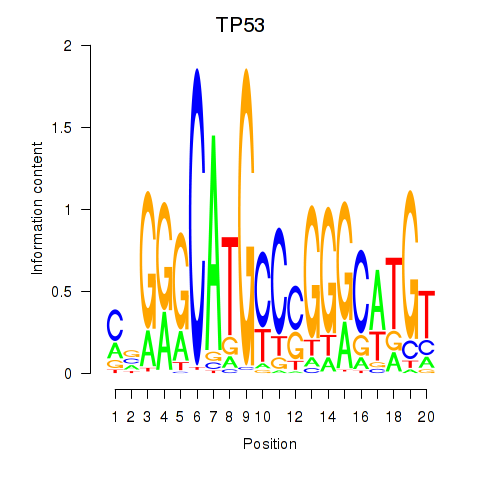
|
| TP63 | 0.336 | 0.985 |
|
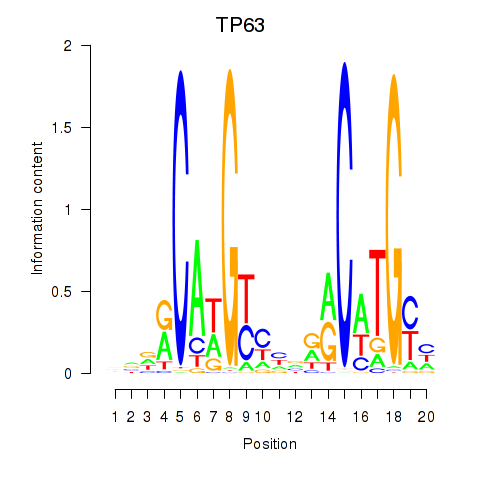
|
| TP73 | -0.306 | -0.456 |
|
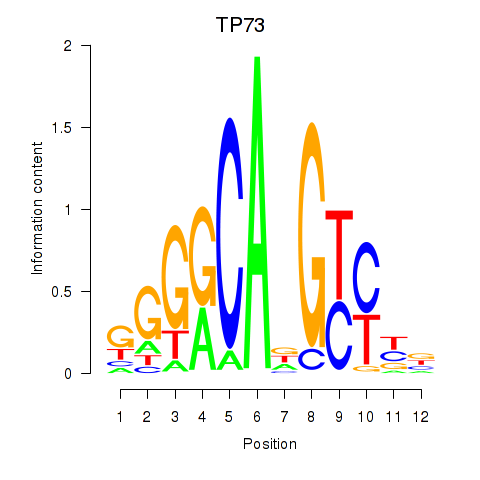
|
| TWIST1_SNAI1 | -0.056 | -0.250 |
|
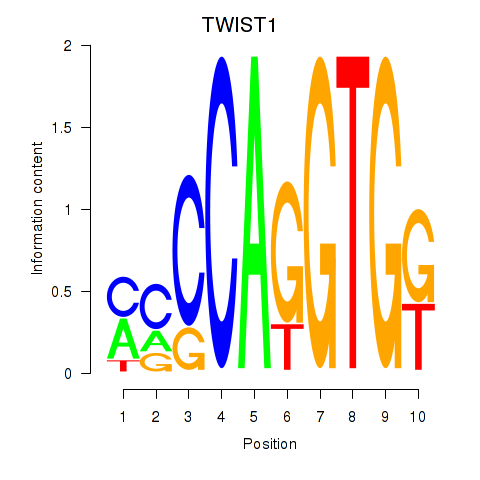
|
| UAAGACG | 1.693 | 0.646 |
|

|
| UAAGACU | -1.347 | -0.481 |
|

|
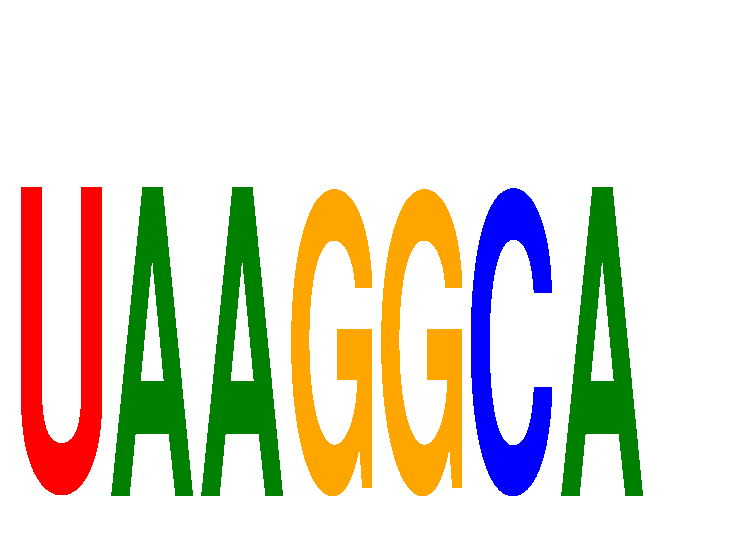
| UAAGGCA | 0.336 | 0.905 |
|

|
| UAAUGCU | -0.563 | -0.662 |
|

|
| UACAGUA | 0.713 | 0.803 |
|

|
| UAGUGUU | 0.577 | 0.977 |
|
|
| UAUUGCU | -0.159 | -0.513 |
|

|
| UBP1 | 0.227 | 0.281 |
|
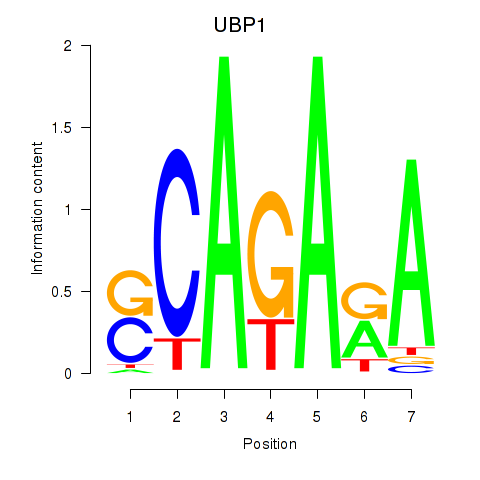
|
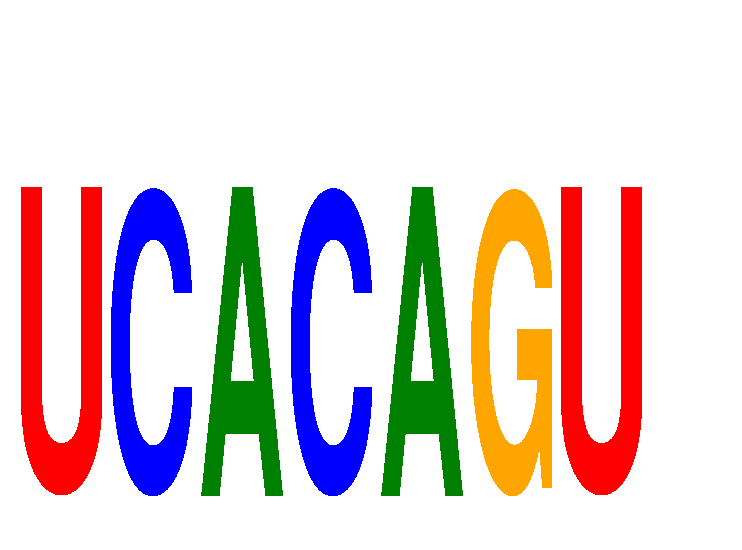
| UCACAGU | -0.269 | -0.539 |
|

|
| UCACAUU | -0.145 | -0.246 |
|

|
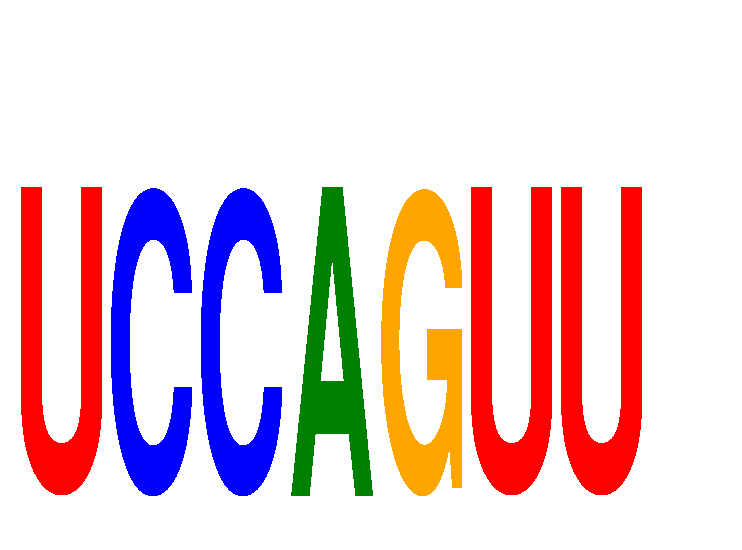
| UCCAGUU | 0.430 | 1.007 |
|

|
| UCCCUUU | 0.331 | 0.304 |
|
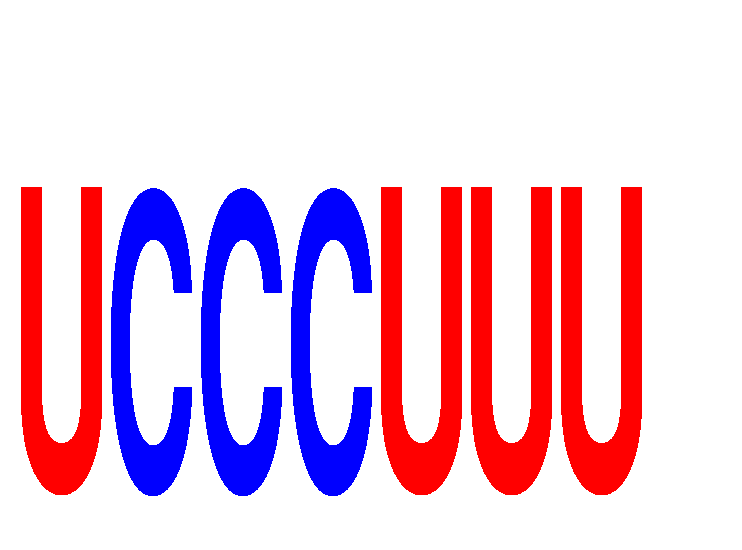

|
| UGAAAUG | 0.442 | 0.503 |
|
|
| UGACAUC | 0.432 | 0.203 |
|

|
| UGACCUA | 0.767 | 0.314 |
|

|
| UGCAGUC | 0.039 | 0.033 |
|

|
| UGCAUAG | -0.066 | -0.178 |
|

|
| UGCAUUG | 0.496 | 0.416 |
|

|
| UGGCACU | 0.437 | 0.809 |
|
|
| UGGUCCC | 0.106 | 0.211 |
|

|
| UGUGCGU | 2.318 | 0.488 |
|

|
| UGUGCUU | -0.308 | -0.861 |
|

|
| UUGGCAA | 0.503 | 0.861 |
|

|
| UUGGCAC | -0.731 | -1.222 |
|

|
| UUGGUCC | 0.101 | 0.210 |
|

|
| UUGUUCG | -0.116 | -0.103 |
|

|
| UUUUUGC | -0.896 | -0.180 |
|

|
| VAX1_GSX2 | -0.485 | -0.804 |
|
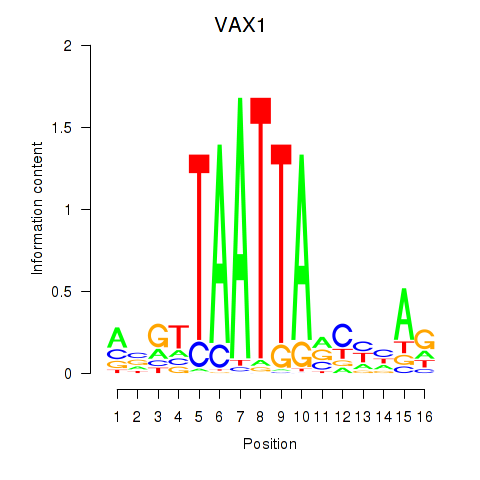
|
| VAX2_RHOXF2 | -0.130 | -0.152 |
|
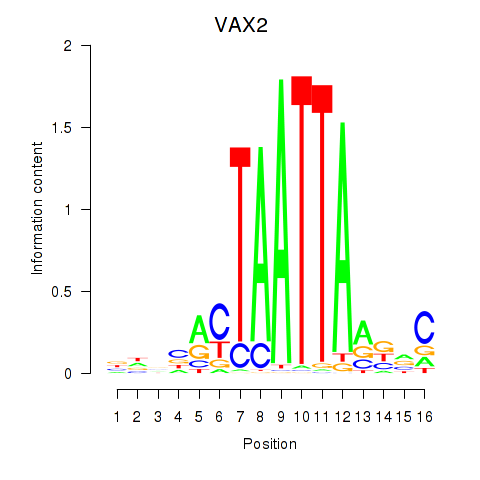
|
| VDR | -0.049 | -0.138 |
|
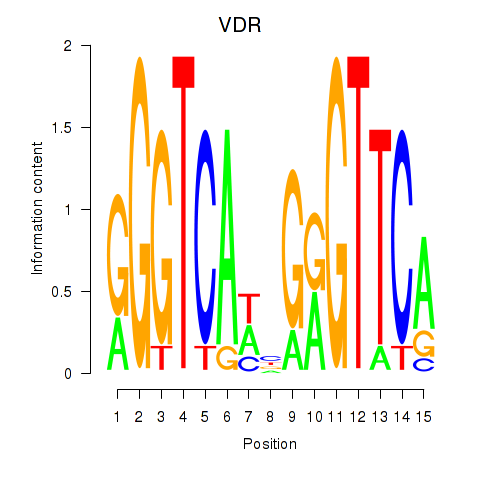
|
| VENTX | 0.019 | 0.031 |
|
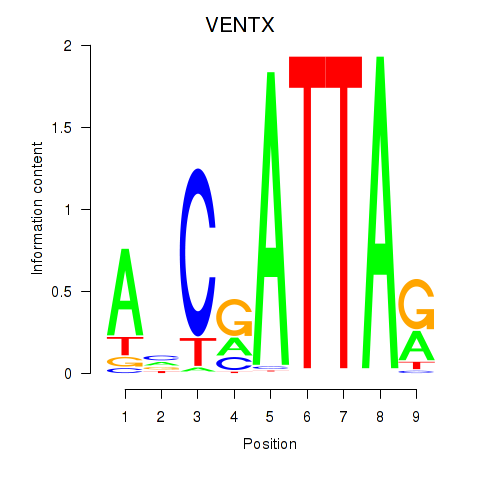
|
| VSX1 | 0.296 | 0.747 |
|
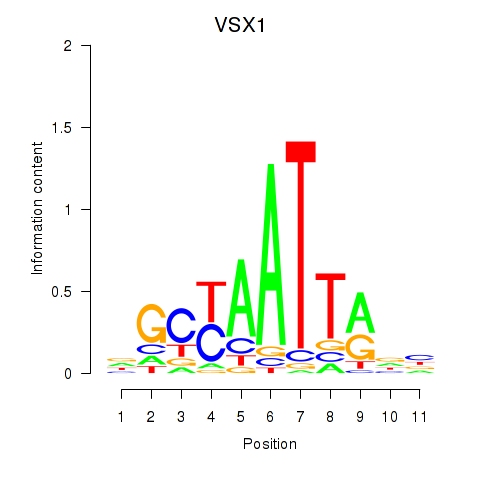
|
| WRNIP1 | -0.124 | -2.289 |
|
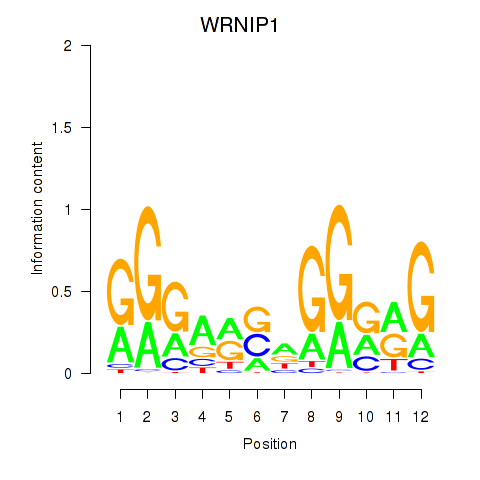
|
| WT1_MTF1_ZBTB7B | -0.068 | -1.102 |
|
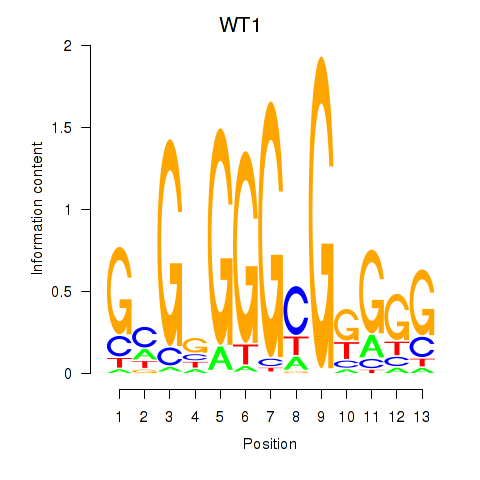
|
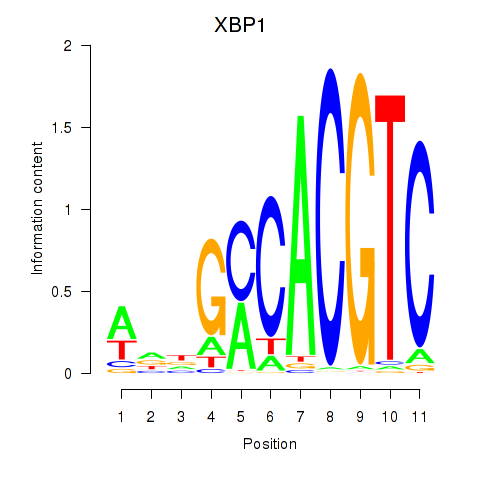
| XBP1 | 0.451 | 1.012 |
|
|
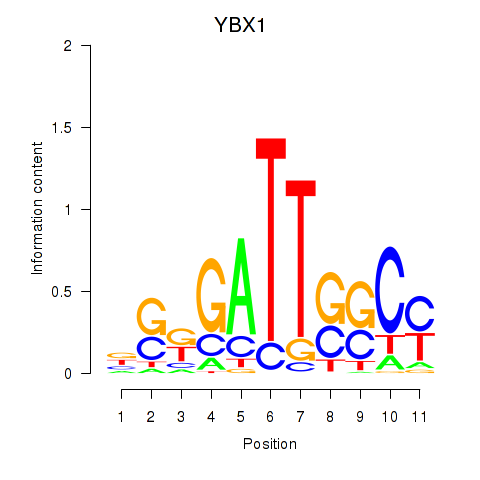
| YBX1_FOS_NFYC_NFYA_NFYB_CEBPZ | 0.132 | 2.626 |
|
|
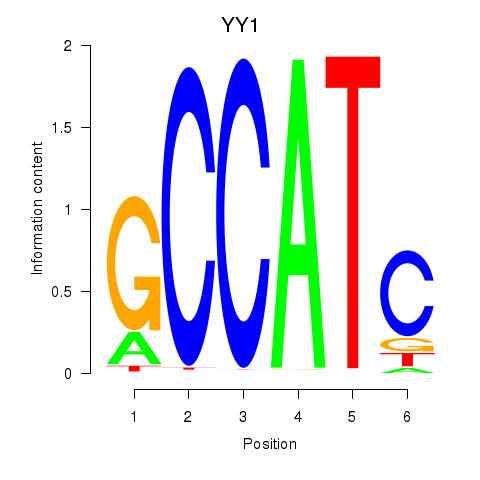
| YY1_YY2 | 0.259 | 1.373 |
|
|
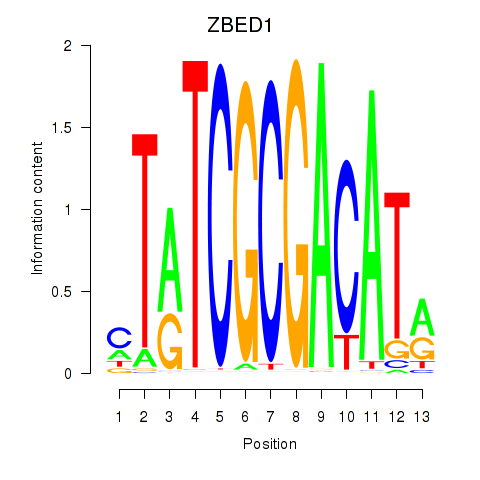
| ZBED1 | -1.500 | -1.027 |
|
|
| ZBTB12 | 0.272 | 0.465 |
|
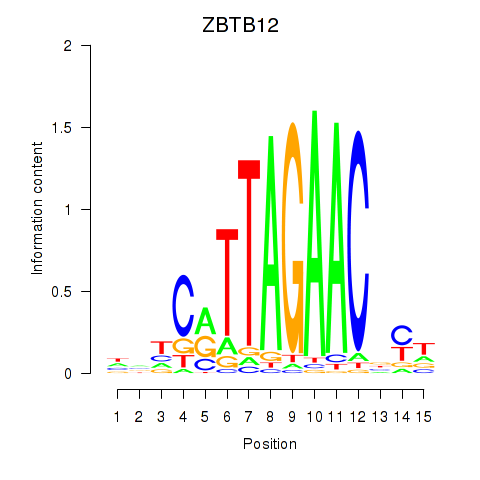
|
| ZBTB14 | -0.177 | -1.215 |
|
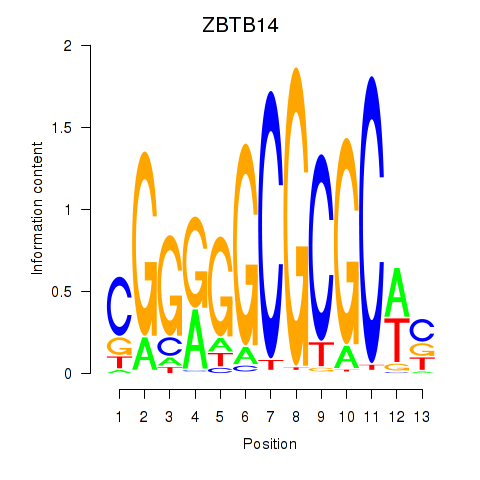
|
| ZBTB16 | 0.146 | 0.358 |
|
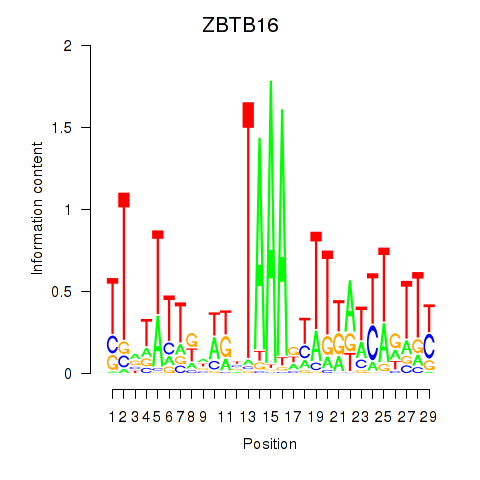
|
| ZBTB18 | -0.201 | -0.435 |
|
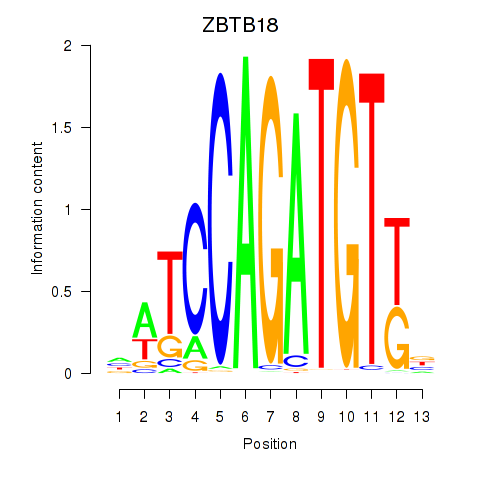
|
| ZBTB3 | -0.114 | -0.174 |
|
|
| ZBTB33_CHD2 | 0.538 | 3.775 |
|
|
| ZBTB4 | 0.014 | 0.030 |
|
|
| ZBTB49 | 0.005 | 0.007 |
|
|
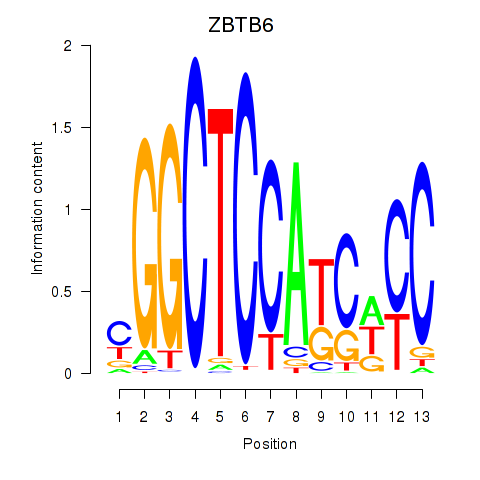
| ZBTB6 | -0.201 | -0.986 |
|
|
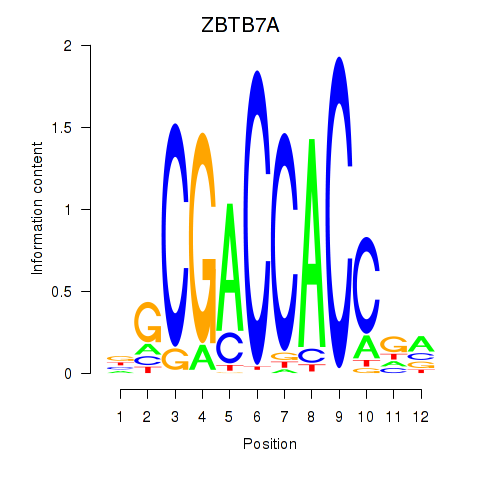
| ZBTB7A_ZBTB7C | 0.095 | 0.218 |
|
|
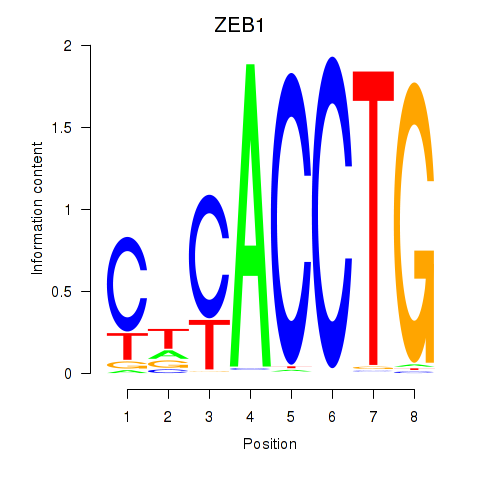
| ZEB1 | -0.126 | -1.295 |
|
|
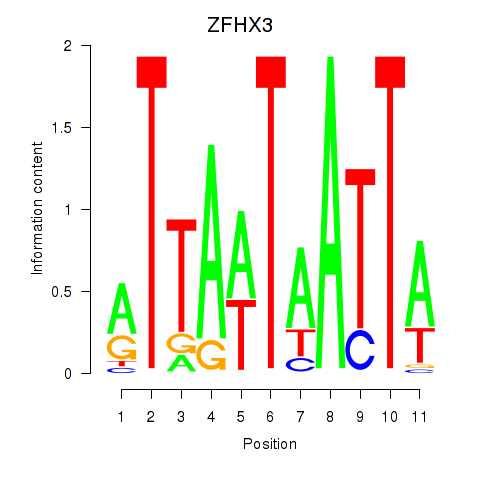
| ZFHX3 | -0.187 | -0.226 |
|
|
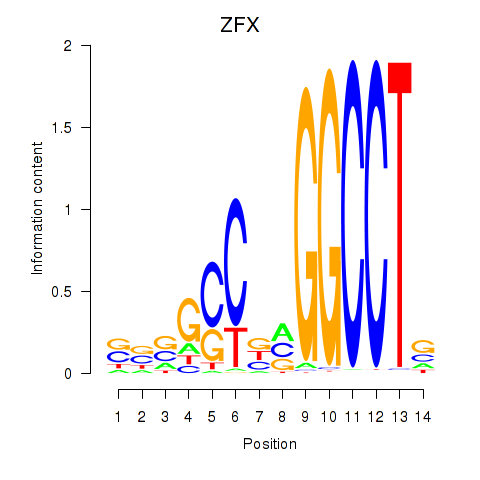
| ZFX | 0.158 | 1.274 |
|
|
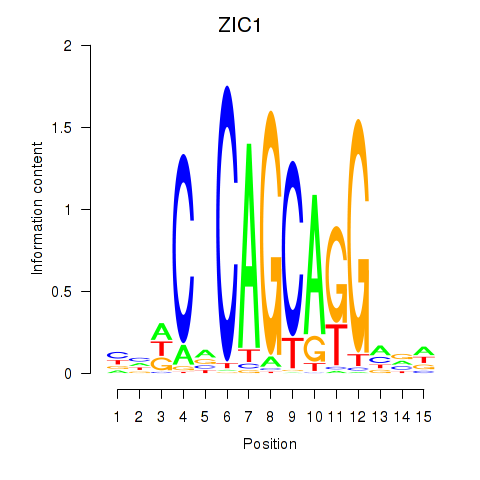
| ZIC1 | -0.446 | -0.914 |
|
|
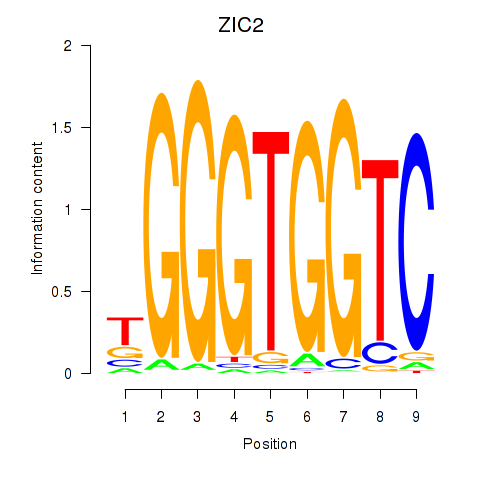
| ZIC2_GLI1 | -0.381 | -0.943 |
|
|
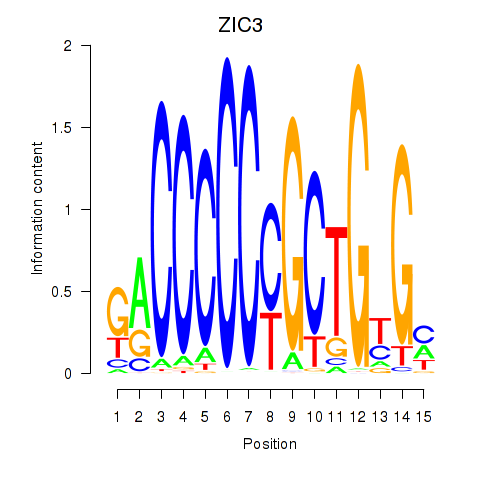
| ZIC3_ZIC4 | -0.152 | -0.558 |
|
|
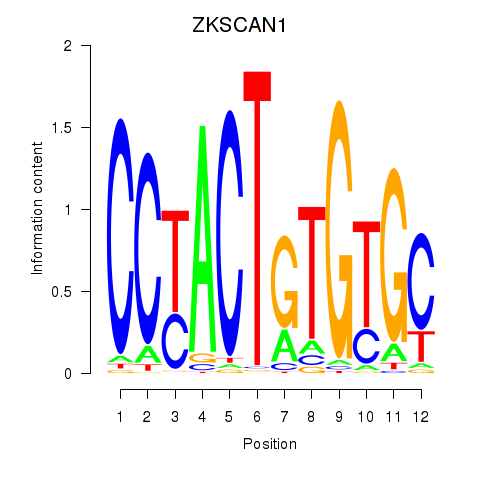
| ZKSCAN1 | 0.138 | 0.280 |
|
|
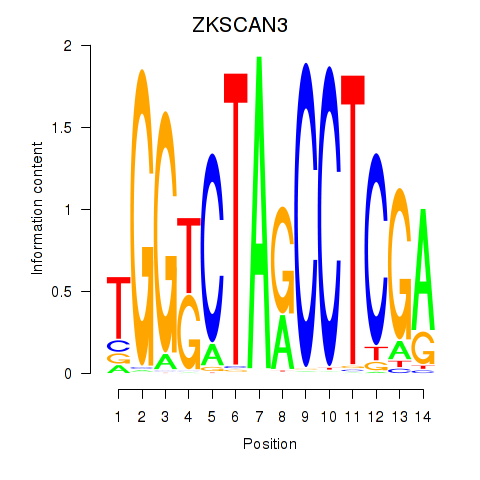
| ZKSCAN3 | 0.159 | 0.199 |
|
|
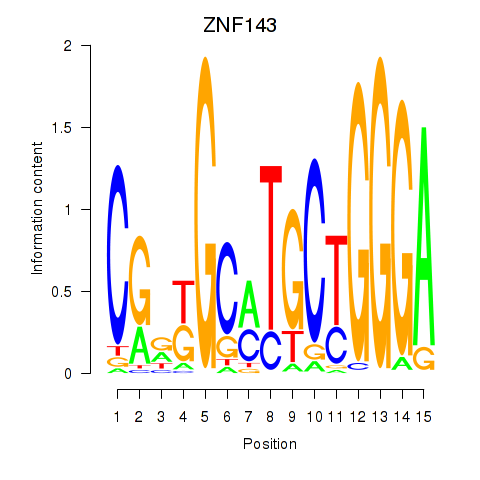
| ZNF143 | 0.218 | 1.202 |
|
|
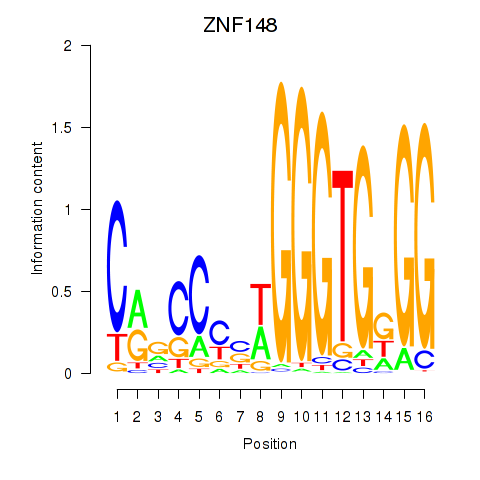
| ZNF148 | 0.013 | 0.050 |
|
|
| ZNF232 | 0.065 | 0.080 |
|
|
| ZNF263 | -0.177 | -1.139 |
|
|
| ZNF274 | 0.213 | 0.221 |
|
|
| ZNF282 | 0.317 | 0.563 |
|
|
| ZNF333 | -0.476 | -0.769 |
|
|
| ZNF35 | -0.843 | -0.781 |
|
|
| ZNF350 | -0.258 | -0.404 |
|
|
| ZNF384 | 0.141 | 1.297 |
|
|
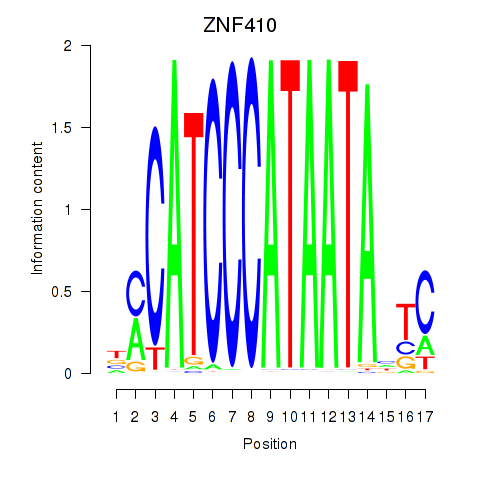
| ZNF410 | 0.378 | 0.482 |
|
|
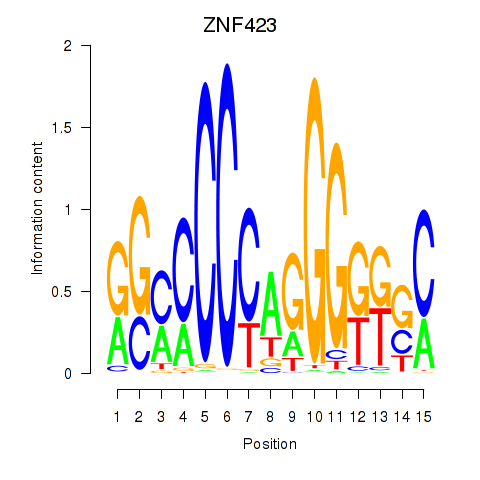
| ZNF423 | -0.087 | -0.161 |
|
|
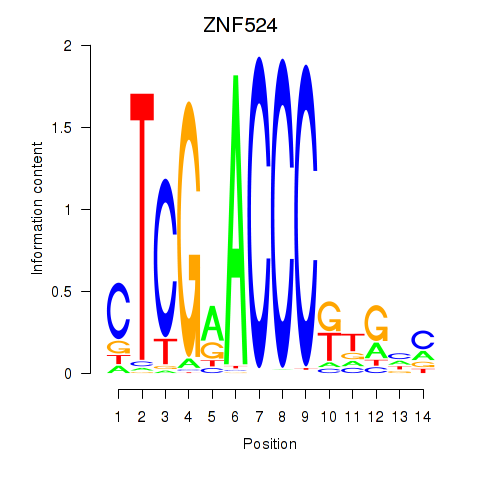
| ZNF524 | 0.010 | 0.064 |
|
|
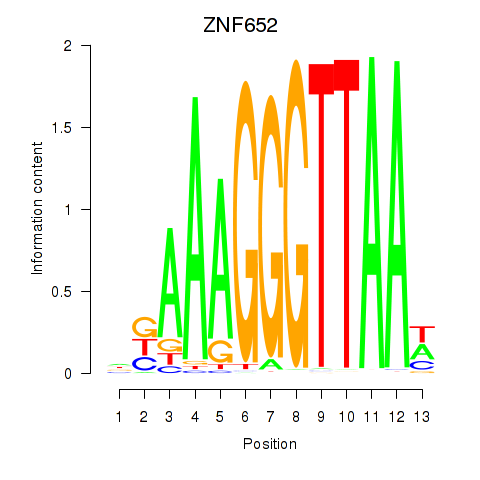
| ZNF652 | -0.003 | -0.011 |
|
|
| ZNF691 | -0.448 | -0.753 |
|
|
| ZNF711_TFAP2A_TFAP2D | 0.115 | 2.830 |
|
|
| ZNF740_ZNF219 | 0.194 | 0.738 |
|
|
| ZNF784 | -0.204 | -0.592 |
|
|
| ZNF8 | 0.738 | 0.906 |
|
|
| ZSCAN4 | -0.019 | -0.039 |
|
|